- The paper demonstrates that PLS-Net achieves an F1-score up to 88% with significantly faster convergence than traditional 3D U-Net.
- It details a robust preprocessing pipeline and mixed precision training using TensorFlow and PyTorch to optimize segmentation.
- The study highlights clinical applicability with rapid inference speeds on both GPU and CPU, enabling near-real-time diagnostics.
Fast Meningioma Segmentation in T1-weighted MRI Volumes Using a Lightweight 3D Deep Learning Architecture
Introduction
This paper addresses the automation of meningioma segmentation within T1-weighted MRI volumes through the employment of lightweight 3D deep learning architectures. The study focuses on two distinct neural network architectures: an encoder-decoder model akin to a 3D U-Net and a more efficient multi-scale alternative called PLS-Net. By leveraging datasets featuring both surgically treated and conservatively managed meningiomas, the study targets improvements in segmentation accuracy and processing speed. The overarching aim is to facilitate clinical applications by reducing time investment and increasing reliability.
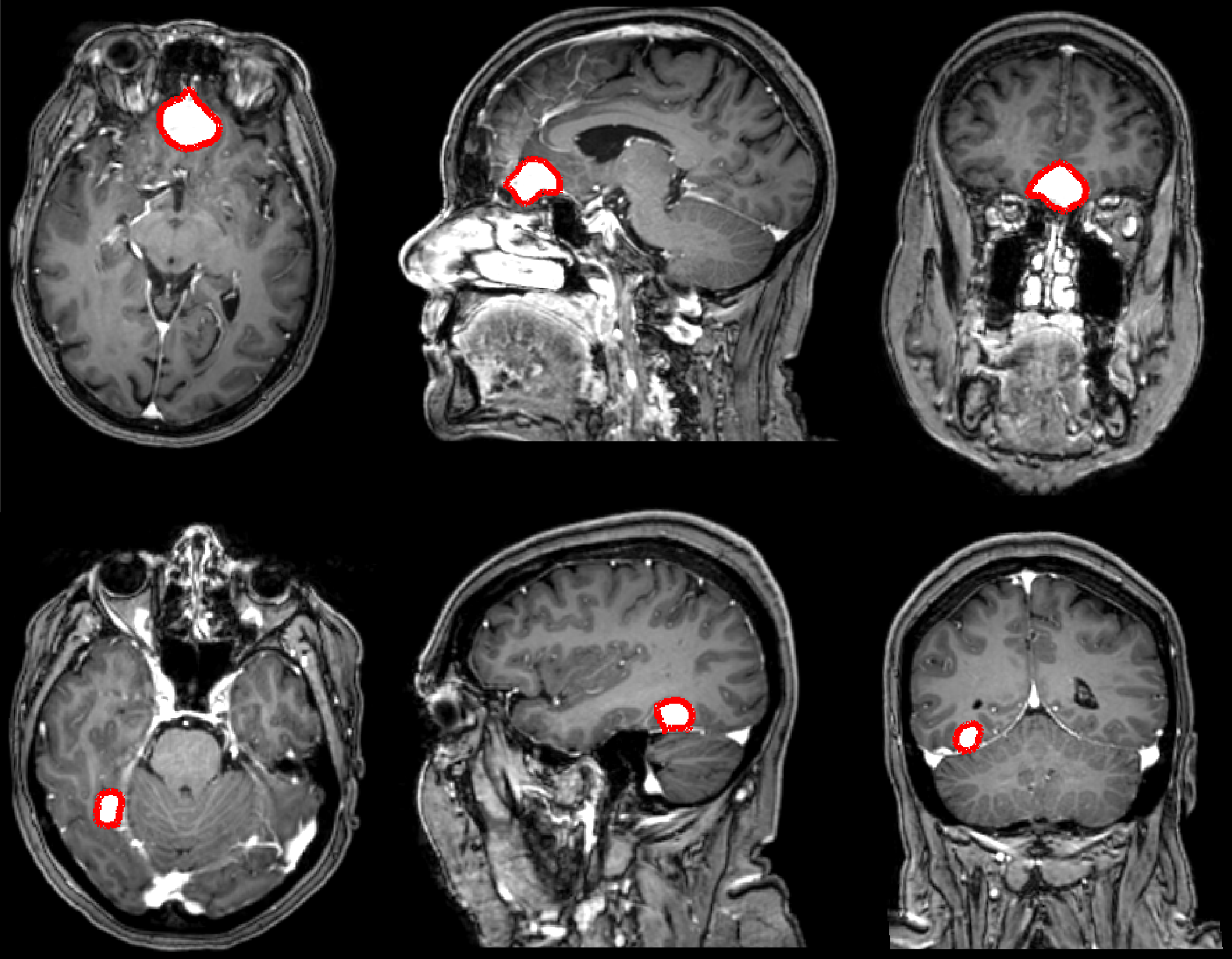
Figure 1: Illustrations of the manually annotated meningiomas over the dataset (in red), showing different views per patient.
Methodology
Data Representation and Preprocessing
The dataset comprises 698 T1-weighted MRI volumes sourced from diverse institutions, ensuring variability in scanning conditions and reducing bias. Preprocessing steps include N4 bias correction, resampling to isotropic spacing, volume cropping, and standardized intensity normalization. Volumes were stratified into subsets based on image quality, allowing segregated model training strategies.
Neural Network Architectures
3D U-Net
The 3D U-Net employed in this study consists of several convolutional layers designed to capture volumetric context. Input volumes undergo slab-based division due to memory constraints, a process that potentially impairs global structural understanding.
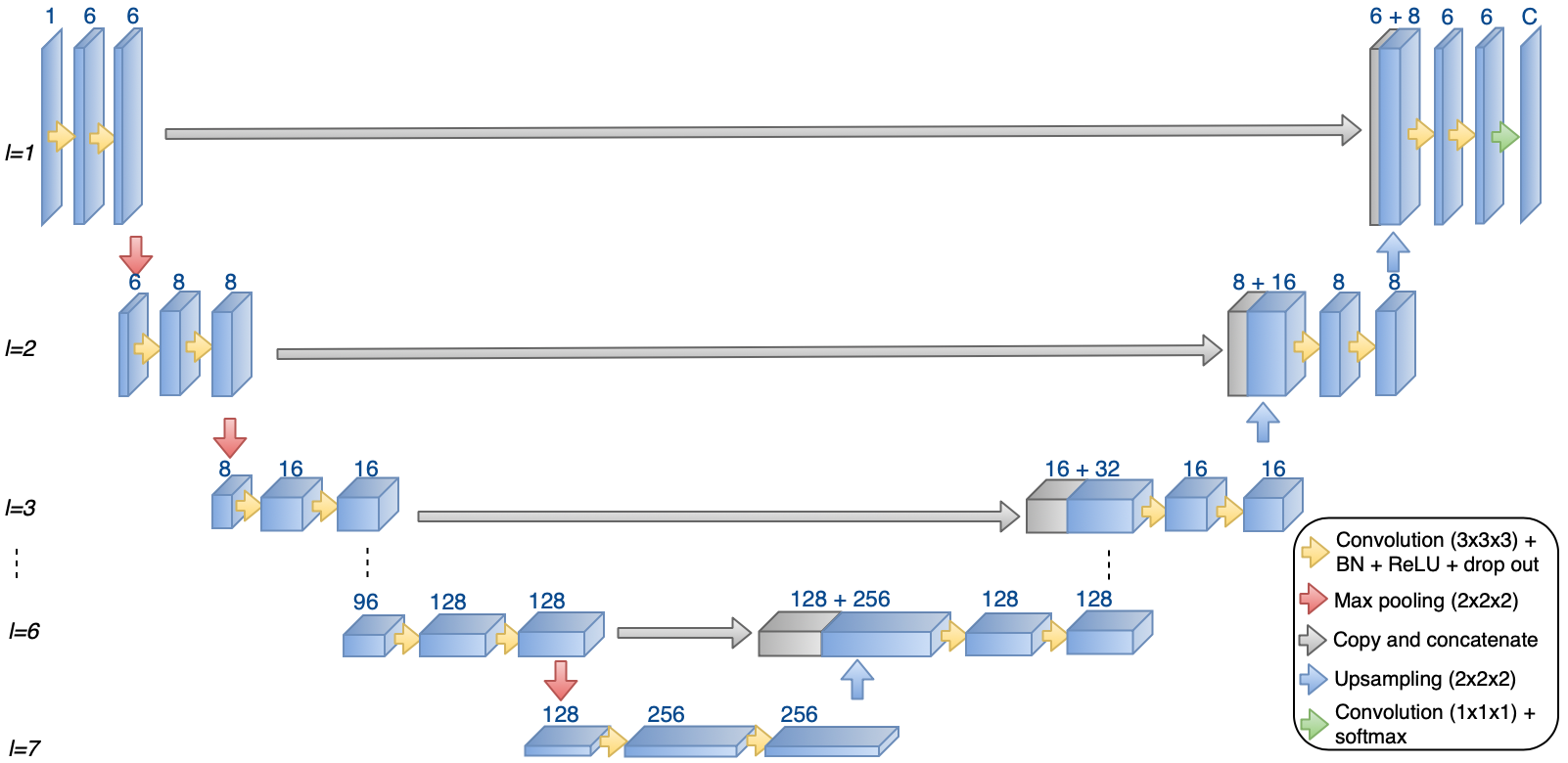
Figure 2: 3D U-Net architecture used in this study; adaptable in terms of depth and layer configurations according to input resolution.
PLS-Net
PLS-Net improves upon traditional models through multi-scale feature processing and depthwise separable convolutions. This architecture ensures high-resolution input handling and significantly reduces computational demands.
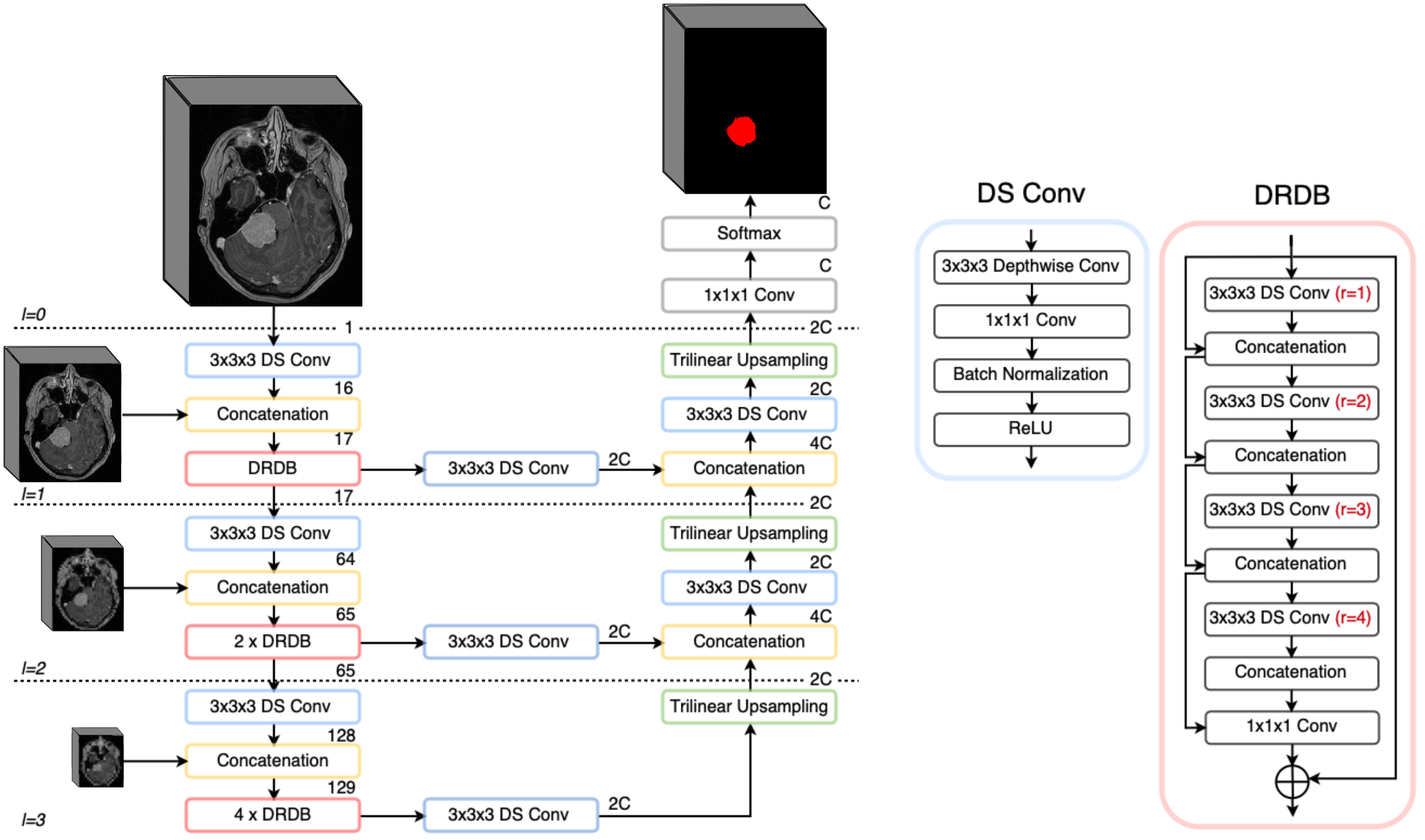
Figure 3: PLS-Net architecture utilized, highlighting its streamlined nature tailored for efficient segmentation tasks.
Training Protocols
Training occurred via contrasting frameworks: TensorFlow for full precision and PyTorch for mixed precision, maximizing computational efficiency. Augmentation tactics encompassed spatial transformations to bolster model robustness. Mixed precision training leveraged reduced memory consumption, allowing substantial computational gain without compromising model precision.
Results
PLS-Net demonstrated dominant performance, achieving an F1-score of up to 88%, noticeably exceeding U-Net's capabilities. Speed evaluations revealed that PLS-Net models converged rapidly, in under 50 hours, while U-Net necessitated 130 hours, underscoring PLS-Net's operational advantages.
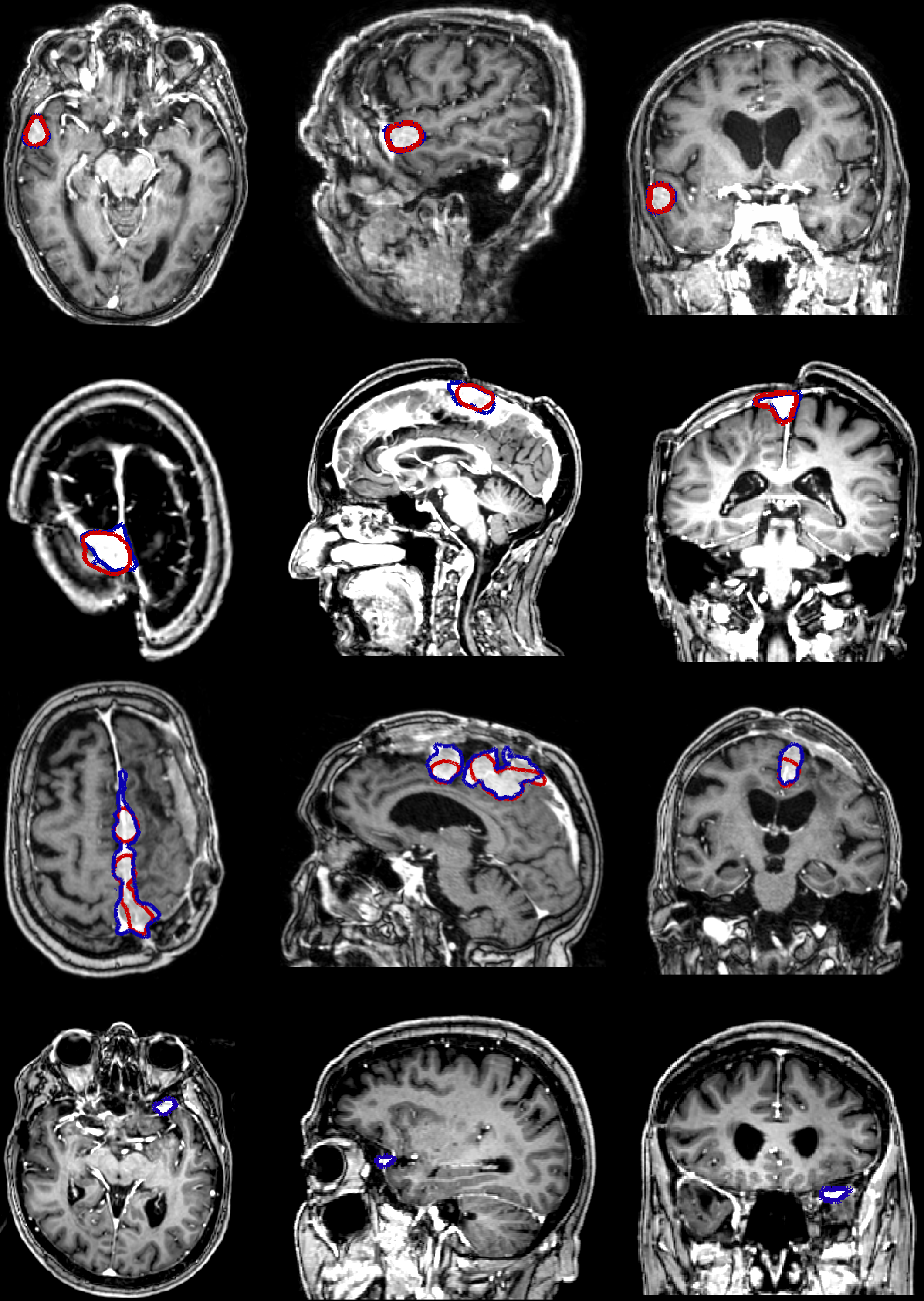
Figure 4: Illustrations of segmentation results using the PLS-cfg4 model.
Processing Efficiency
On conventional hardware, PLS-Net demonstrated swift inference speeds, completing on GPU within a second and on CPU within 15 seconds. This rapid inference aligns well with clinical demand for near-real-time image processing.
Image Resolution Impact
Performance on high-resolution images outperformed low-resolution counterparts, suggesting a correlation between input detail and segmentation accuracy. However, the algorithm demonstrated resilience to image degradation, maintaining essential performance levels across resolutions.
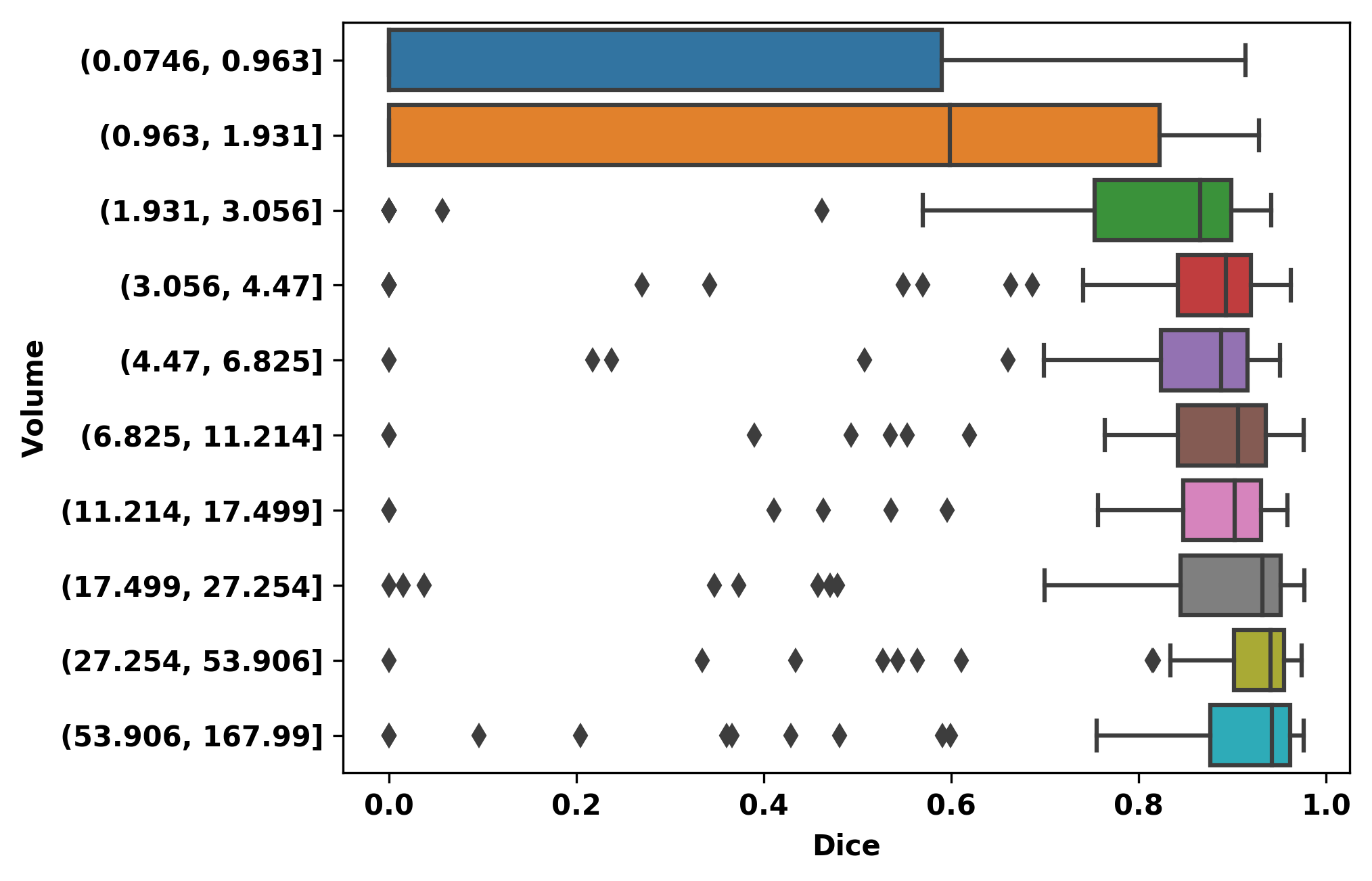
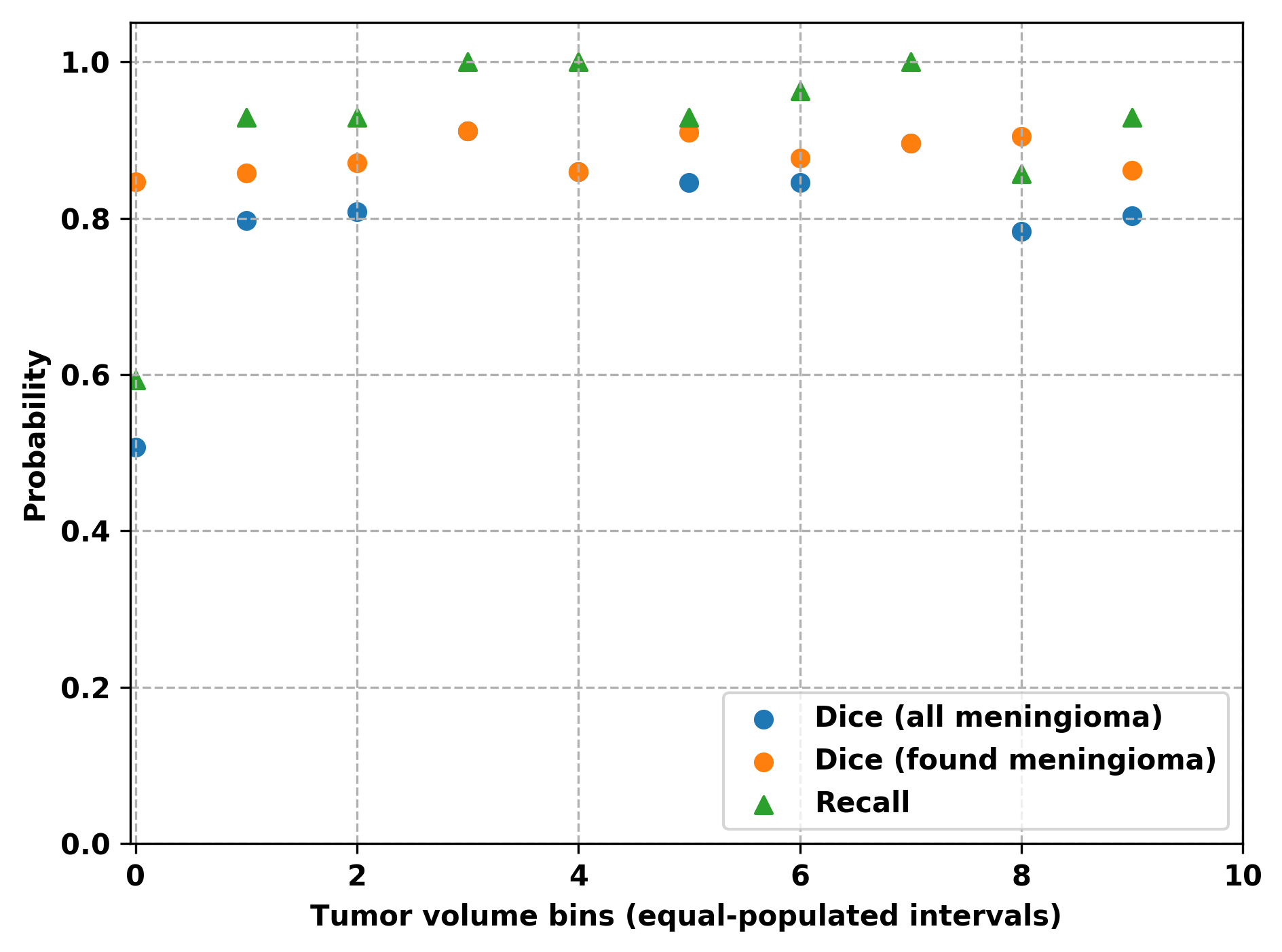
Figure 5: Overall, hospital, and outpatient clinic results for PLS-cfg4, illustrating the segmentation performance in terms of Dice and F1-scores over varying tumor volumes.
Discussion
The effectiveness of the lightweight PLS-Net architecture is notable, meriting its adoption in settings constrained by computational resources. The demonstrated robustness of segmentation with limited input resolution fluctuations highlights its potential for widespread clinical application. Future research should focus on refining the detection of small meningiomas, potentially through enhanced input resolution and more varied data sampling.
Conclusion
This study advances meningioma segmentation through the employment of the PLS-Net architecture, emphasizing clinical applicability through efficient processing. The model's competitive accuracy and reduced computational burdens are promising for adoption in routine clinical workflows, presenting a compelling case for lightweight architectures in medical imaging.
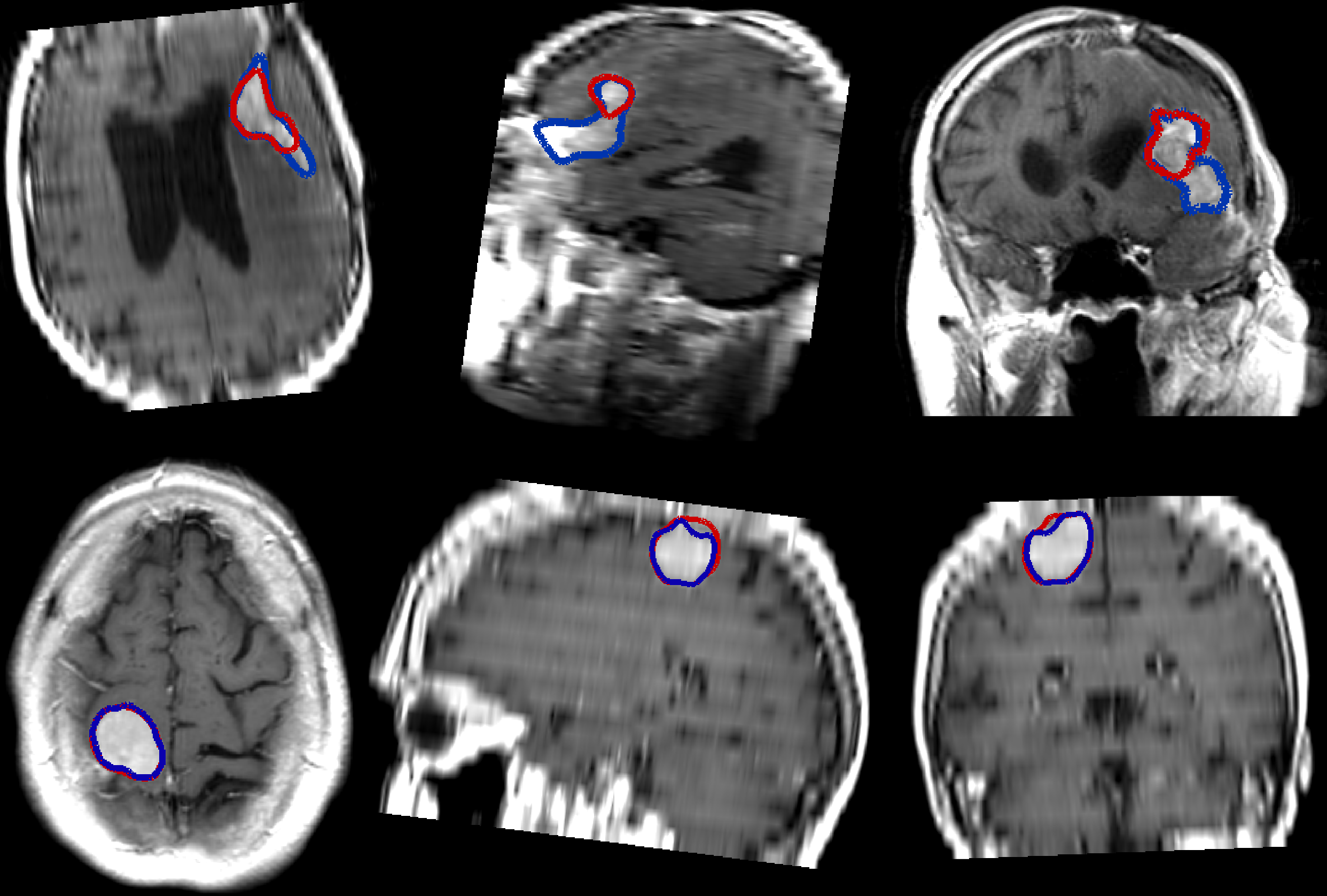
Figure 6: Illustration of segmentation results on images with poor resolution using the PLS-cfg4 model.
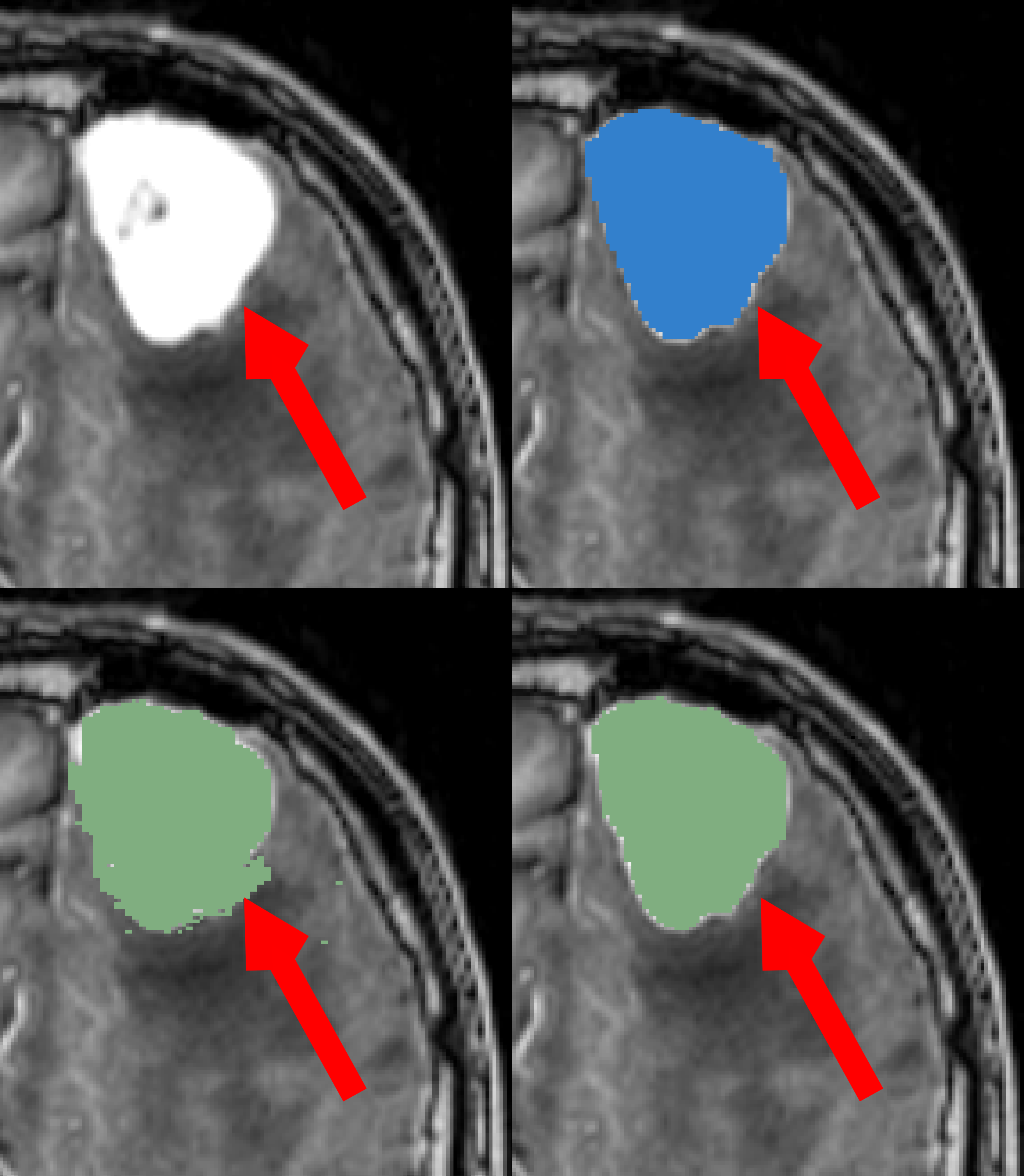
Figure 7: Illustration of ground truth noise, denoting the necessity for further refinement in annotation processes.







