- The paper presents SSRGNet, a unified model merging transformer-based sequence encoding with 3D relation-aware graph networks for improved protein secondary structure prediction.
- It leverages parallel fusion of sequence and spatial features, achieving notable gains in F1-scores on benchmarks like NetSurfP-2.0, CB513, TS115, and CASP12.
- Explicit 3D structural encoding enhances prediction of complex protein motifs and paves the way for more comprehensive functional annotation.
Introduction
Accurate protein secondary structure prediction (PSSP) is foundational for computational biology, as secondary structures inform tertiary structure and functional annotation. Traditional approaches exploit sequence or evolutionary features, often neglecting explicit 3D structural information. This paper proposes SSRGNet, a unified architecture combining a pre-trained transformer-based protein LLM (DistilProtBert) for sequence encoding, and relational Graph Convolutional Networks (R-GCN) for context-aware message passing over residue graphs, thereby modeling both sequential and spatial relationships among amino acids.
Model Architecture
The SSRGNet design employs a dual-encoder scheme:
- Sequence Encoder: Utilizes DistilProtBert, a compressed version of ProtBert-UniRef100, to extract context-aware embeddings via masked language modeling, capturing evolutionary and local sequence contexts.
- Graph Encoder: Constructs a multi-relational graph representation from the protein's 3D coordinates. Nodes represent residues (via their α-carbon), while edges encode three key relations: sequential proximity, Euclidean spatial proximity (<10 Å), and nearest-neighborhood (kn=10).
Within this framework, the R-GCN aggregates local and nonlocal structural information by relation-specific transformations, enabling explicit modeling of residue-residue interactions.
Fusion Strategies and Ablation
Features from both encoders are projected to a unified dimension and fused via three strategies: series (element-wise addition), parallel (concatenation), and cross (multi-head attention). Parallel fusion, shown empirically to be most effective, preserves complementary sequence and structure information without explicit prioritization or information loss.
Experimental Setup
All evaluations employ the NetSurfP-2.0 training set with secondary structure classification in both 3-state (H/E/C) and 8-state (B/E/G/H/I/T/S/C) domains. Test sets include CB513, TS115, and CASP12. Datasets are filtered for redundancy and annotated with PDB-derived 3D coordinates.
Performance metrics are Q3/Q8 accuracy and macro F1-score. SSRGNet is compared against unimodal baselines (DCRNN, DeepACLSTM, DistilProtBert) and multimodal reference (GCNBLSTM), with rigorous Bayesian hyperparameter optimization and batch-wise model training via diagonal PyG batching.
Results
SSRGNet outperforms all baselines on all test sets, with marked improvements in macro F1-score.
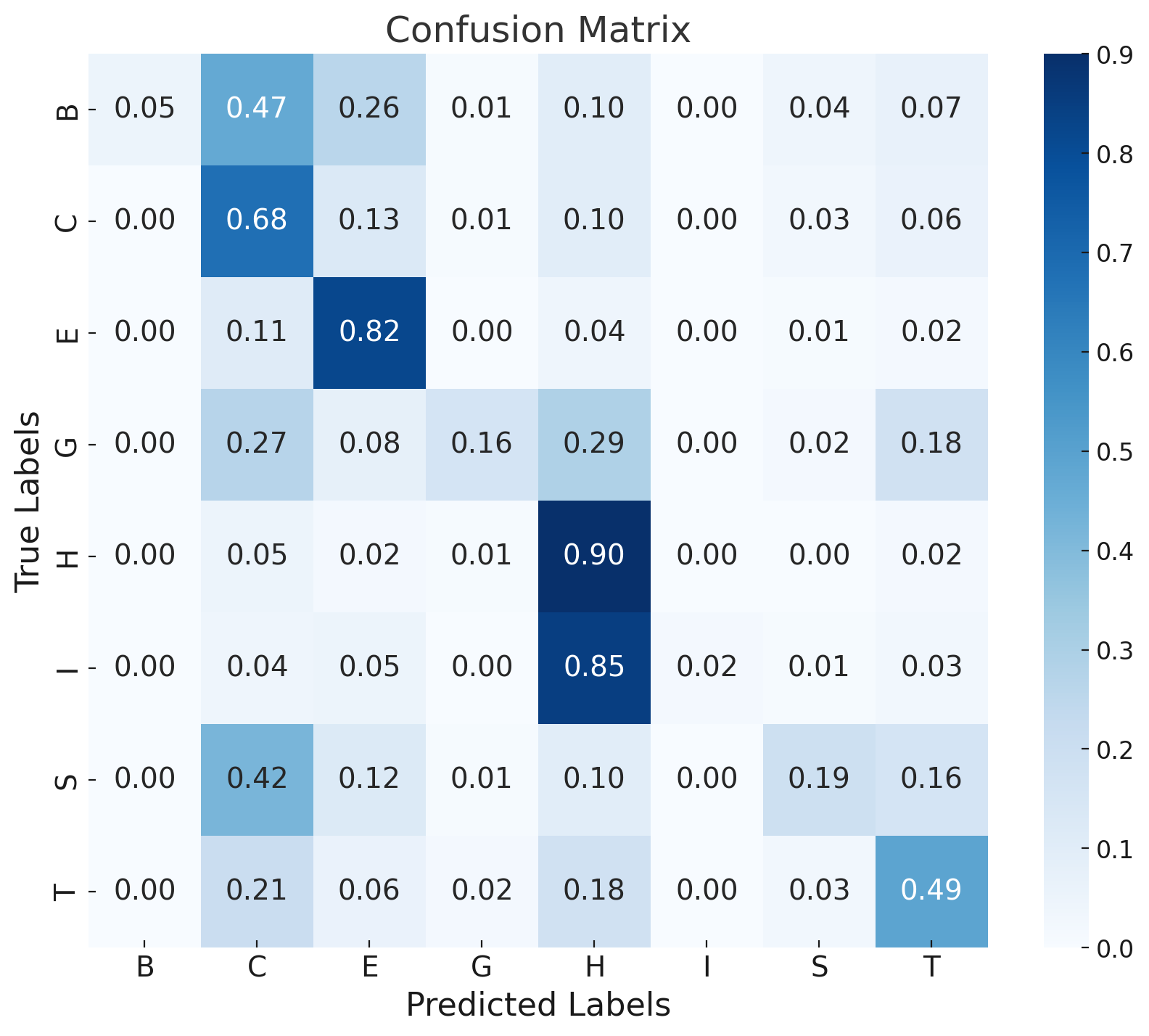
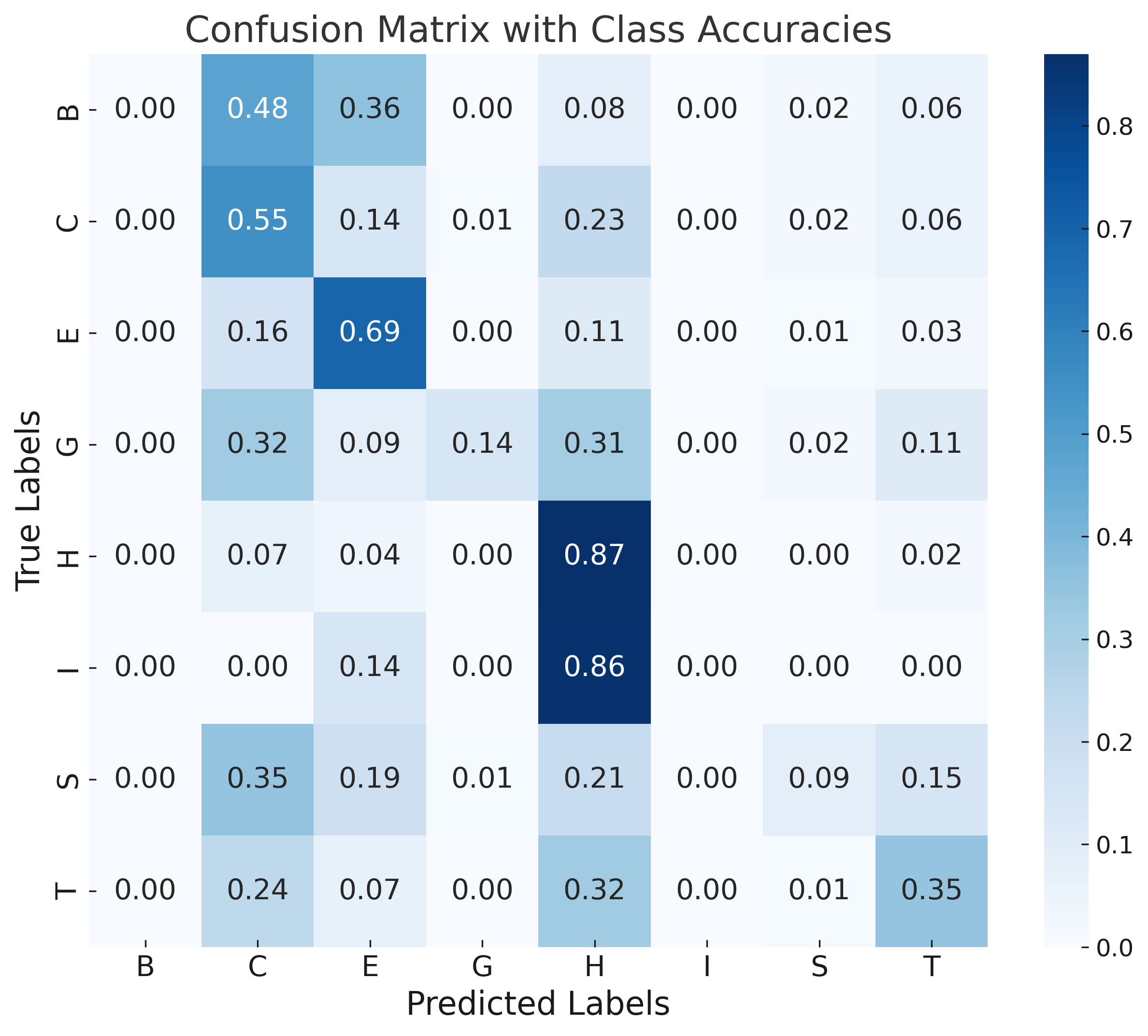
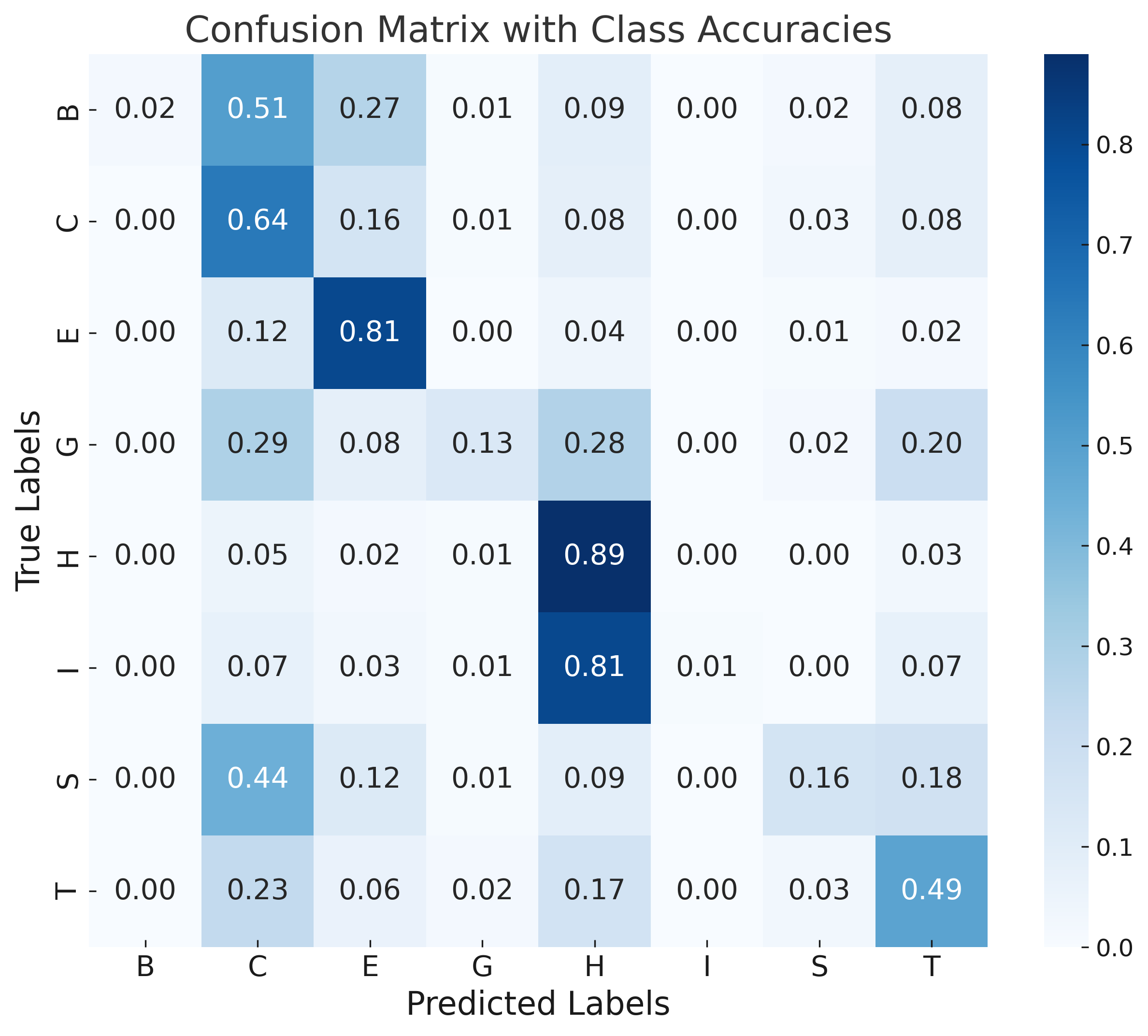
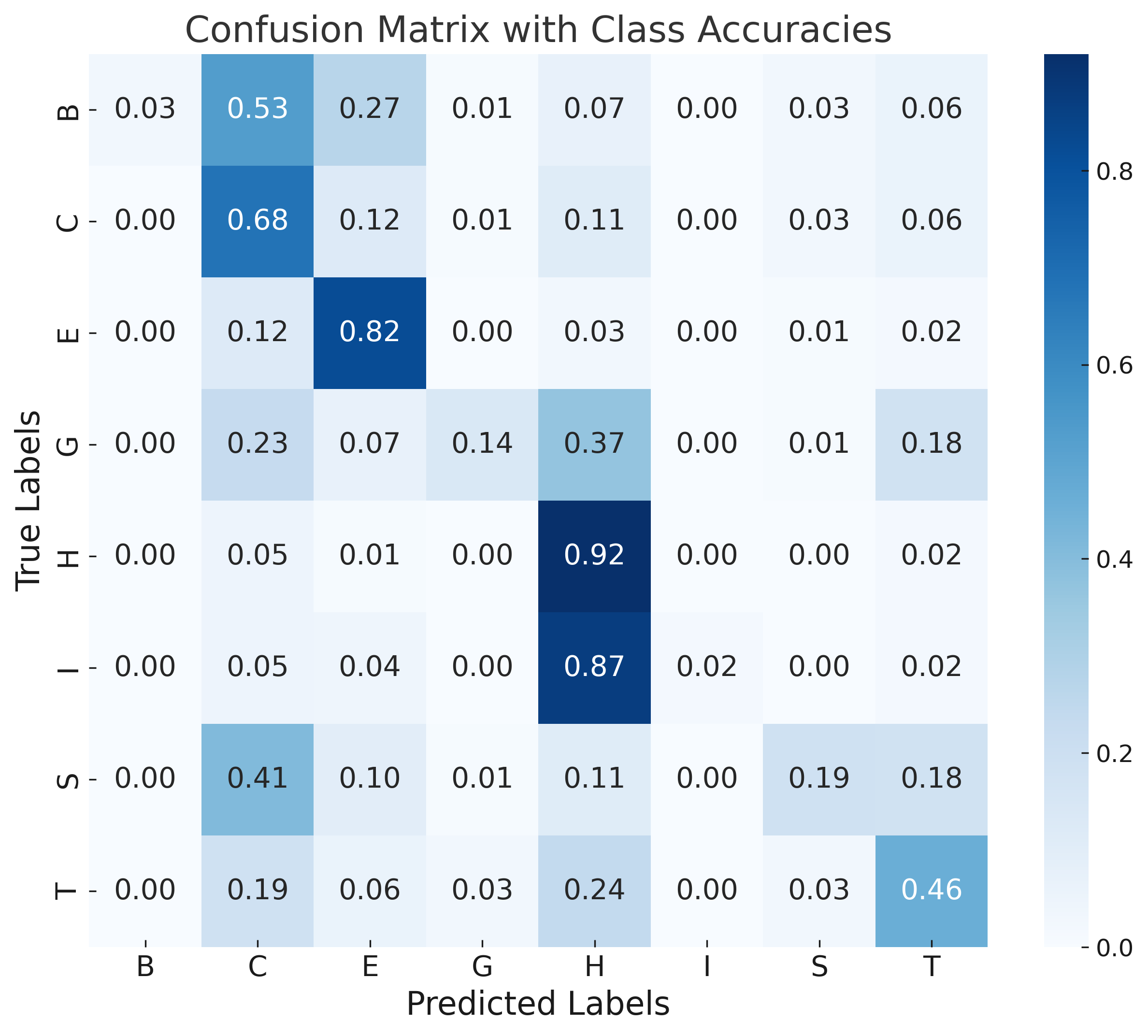
Figure 2: NetSurfP-2.0 benchmark distribution for secondary structure states utilized for training and evaluation.
Notably, SSRGNet achieves F1-scores of 0.67 (3-state, NetSurfP-2.0), 0.61 (CB513), 0.51 (CASP12), and 0.65 (TS115). Compared to vanilla DistilProtBert, this is an absolute gain of ~1–2% across datasets. The improvement is most pronounced in imbalanced and structurally complex protein domains, affirming the contribution of spatial encoding.
Ablation studies reveal that:
- Parallel fusion consistently achieves the best loss, accuracy, and F1 scores versus series or cross fusion.
- Individual relational graphs (R1, R2, R3) each contribute comparably and additively to model performance, confirming the complementary nature of sequential and spatial cues.
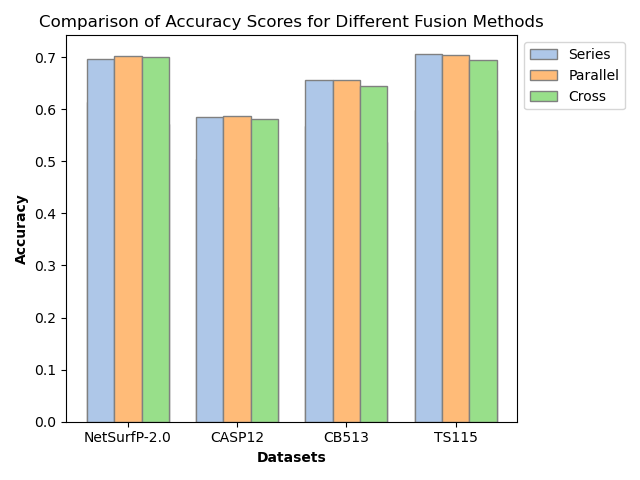
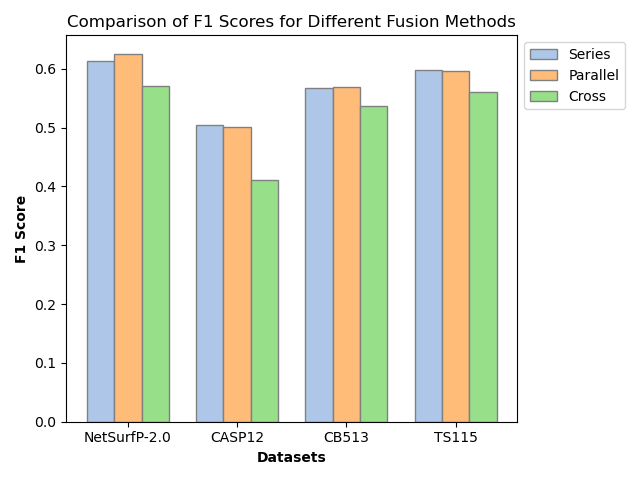
Figure 4: SSRGNet accuracy comparison across fusion strategies and baselines, demonstrating the superiority of parallel fusion and relation-aware graph convolution.
Error analysis via confusion matrices shows rare classes (π-helix, isolated β-bridge) are commonly misclassified due to class imbalance and their structural similarity to more frequent motifs. Sequence-based models (DistilProtBert) dominate overall signal, limiting the marginal gain from structural augmentation under current architectures.
Discussion and Implications
The SSRGNet framework establishes that integrating explicit 3D geometric information via relation-aware message passing substantially enhances PSSP beyond sequence-based deep learning. The modest F1 gains over pre-trained LMs suggest that while sequence context is primary, structural context provides valuable refinement, particularly for less common secondary structure classes and proteins with atypical folding patterns.
Practically, SSRGNet paves the way for deploying efficient, generalized PSSP models capable of handling large-scale sequence repositories alongside PDB structural data. The model's computational complexity is manageable for large protein sets due to efficient batching and fusion; however, further improvements could arise via structure pre-training, attention-based adaptive fusion, and improved graph construction.
Theoretically, explicit relational graph modeling could be extended to contact map prediction, protein design, and function annotation, exploiting advances in geometric deep learning and large protein LMs. SSRGNet's approach is amenable to adaptation for higher-order structure prediction and general molecular property inference.
Conclusion
This work presents SSRGNet, an efficient multi-modal deep learning system fusing transformer-based protein LLMs with relation-aware graph neural networks for explicit spatial encoding. Strong empirical results—particularly in F1-score and accuracy—demonstrate the benefit of integrating 3D structural relations into sequence-based models for protein secondary structure prediction. The research reveals both the promise and current limitations of hybrid architectures in PSSP, serving as a platform for future improvements in sequence-structure integration and more accurate functional annotation in computational biology.





