- The paper introduces a dual-stream network architecture combined with self-supervised contrastive learning to enhance 3D medical image segmentation.
- It employs consistency regularization and pixel-level entropy filtering to mitigate unreliable pseudo-labels in CT and MRI images.
- Experimental results show improved DSC and Jaccard indices on Left Atrial and NIH Pancreas datasets, outperforming previous methods.
Self-Supervised Contrastive Learning for 3D Medical Image Segmentation
This paper introduces a semi-supervised semantic segmentation method designed to leverage both labeled and unlabeled 3D medical imaging data. The core innovation lies in a dual-stream network architecture coupled with a self-supervised contrastive learning paradigm to address the challenges posed by unreliable pseudo-labels. The method demonstrates SOTA results on 3D CT/MRI segmentation datasets.
Methodological Details
The proposed method utilizes a dual-stream network, comprising Subnet A and Subnet B, each with a 3D encoder-decoder architecture to generate prediction maps YA and YB for a given input image. The supervised loss function Ls, as defined in Equation 1, consists of both cross-entropy and Dice losses, guiding the network to learn robust representations for each class based on the labeled data.
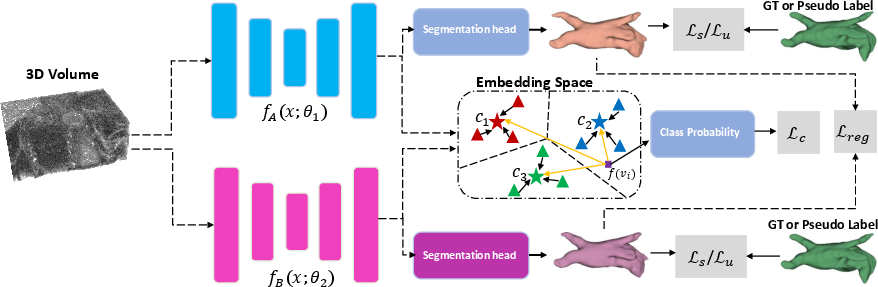
Figure 1: An illustration of the proposed pipeline, highlighting the use of Ls for labeled data and Lu for unlabeled data, with pseudo-labels derived from the network exhibiting lower Ls.
To minimize false predictions, the paper introduces a consistency regularization term, which considers the confidence predictions of one network against the other. For unlabeled images, the method applies a pixel-level entropy-based filtering to exclude unreliable pixel-level pseudo-labels when calculating the unsupervised loss Lu. A regularization loss term, Lreg, is added to Lu to further perform error correction. The overall loss function L combines the supervised loss Ls, unsupervised loss Lu, and a contrastive loss Lc for unreliable pseudo-labels. The weights λu and λc control the contributions of the unsupervised and contrastive losses, respectively.
The consistency regularization refines predictions by identifying voxels where Subnet A and Subnet B produce conflicting predictions despite high confidence. The area of incorrect predictions Mdiff is defined as the set of voxels where the maximum softmax outputs of the two subnetworks differ significantly, exceeding a confidence threshold T. An L1 distance loss function is then applied as a regularization term to correct potential incorrect predictions by each network.
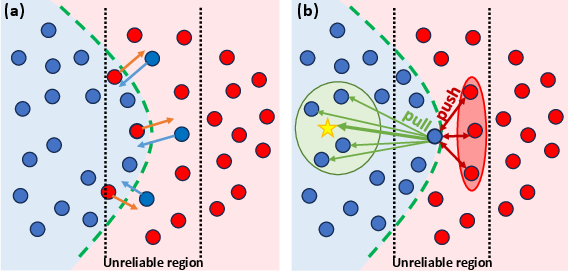
Figure 2: (a) Illustration of the regularization term and (b) contrastive loss effects on prediction refinement, showcasing the method's ability to correct predictions and reduce uncertainty.
The contrastive loss function is designed to mitigate uncertain predictions by guiding low-confidence voxels towards alignment with their corresponding class prototypes. The approach computes the confidence of each voxel's prediction and categorizes them into reliable and unreliable sets. Prototypes for each class are computed as the mean vector of the reliable voxel representations. A distance function is used to compute a distribution over classes for uncertain voxels, and the contrastive loss function aims to move uncertain voxels of the same class towards their respective class prototype while pushing the prototypes of each class away from each other.
Experimental Design and Results
The model was implemented using PyTorch on a single RTX 3090 GPU. The SGD optimizer was used with a weight decay of 0.0001 and a momentum of 0.9. A dynamic learning rate schedule reduced the learning rate by a factor of 10 after every 2500 iterations, for a total of 6000 iterations. The K-fold cross-validation method was used for a robust assessment.
The method was evaluated on the Left Atrial Dataset (LA) and the NIH Pancreas Dataset. The LA dataset consists of 1003 3D gadolinium-enhanced MR imaging volumes. The NIH Pancreas Dataset comprises 82 abdominal CT volumes with manual pancreas annotations.
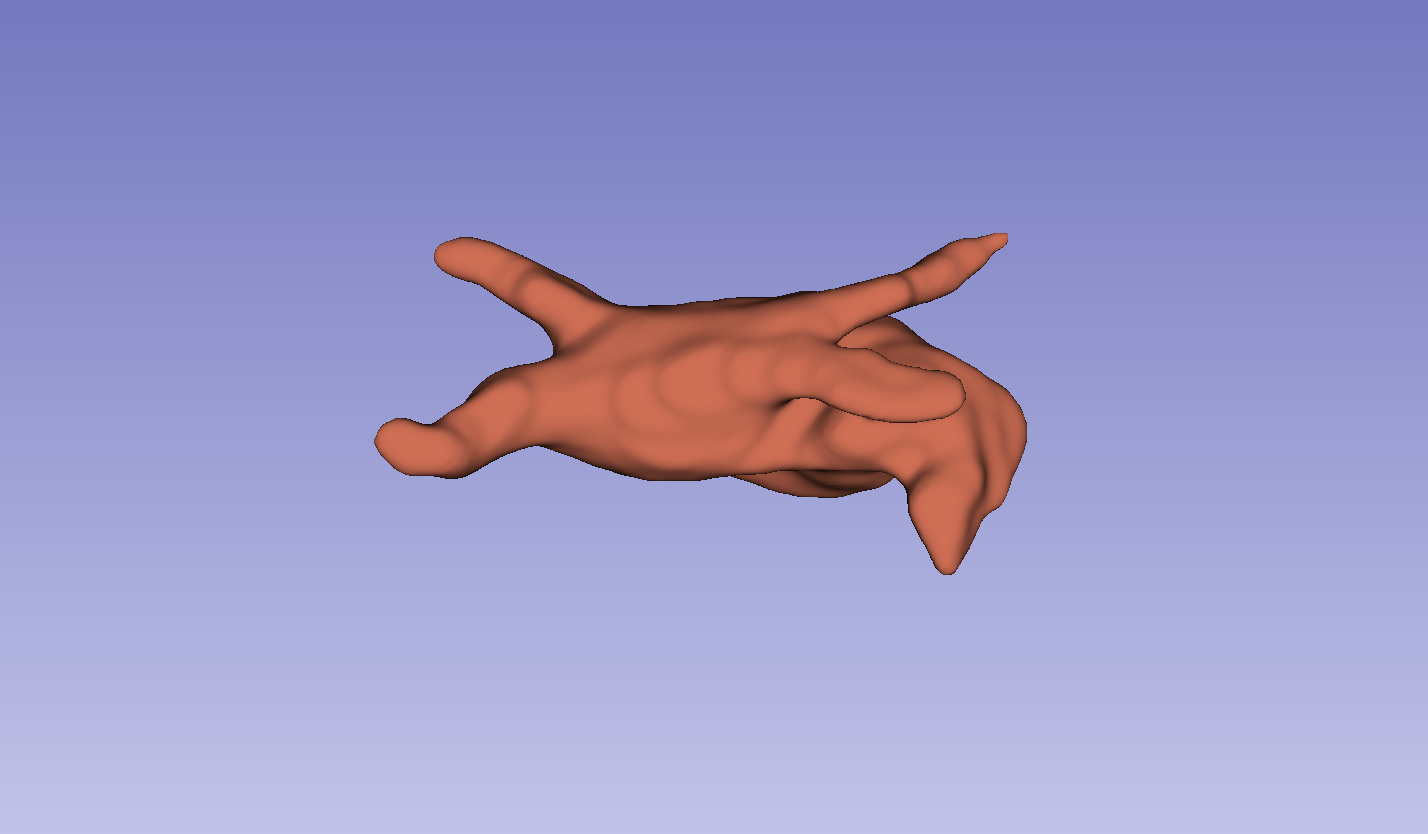
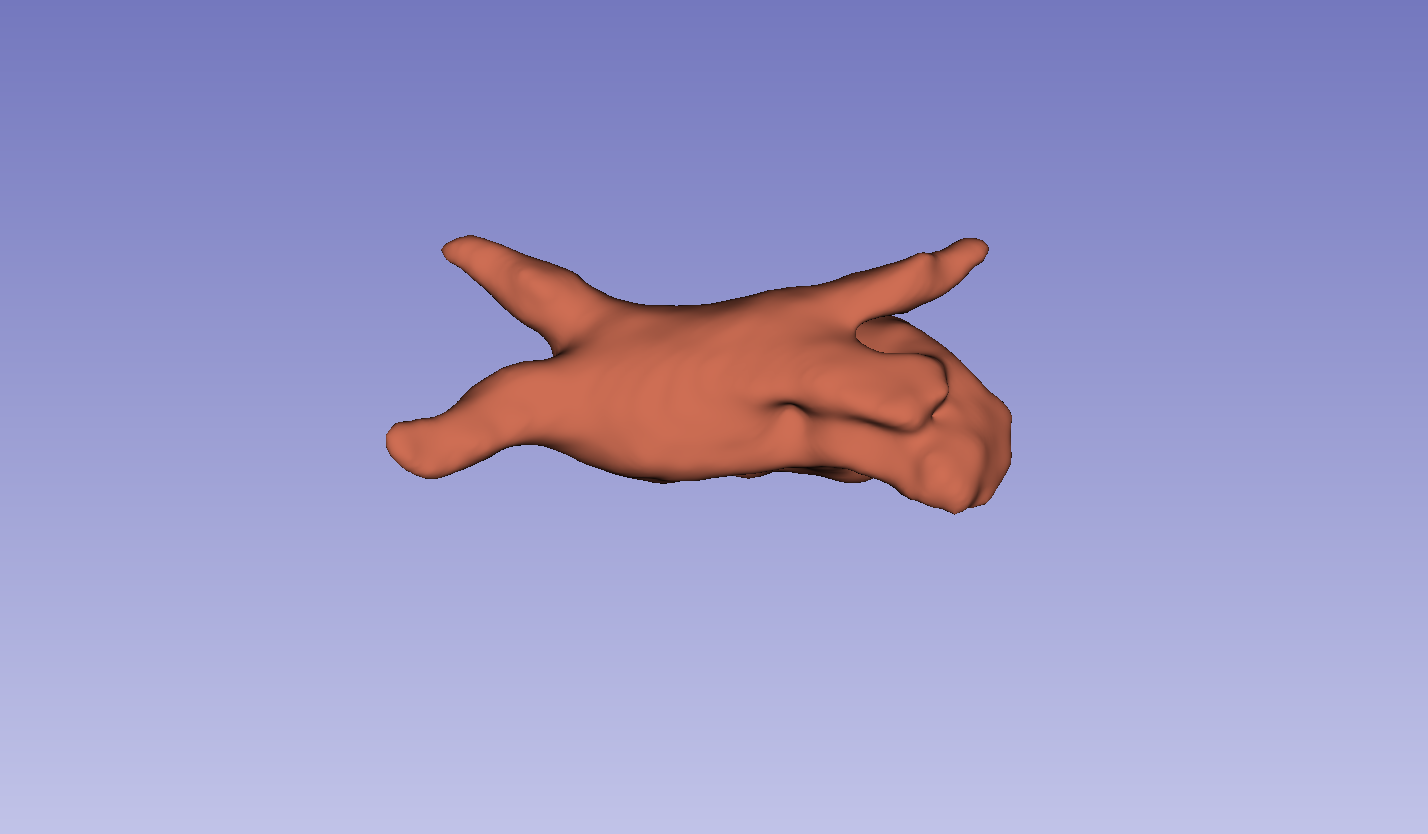
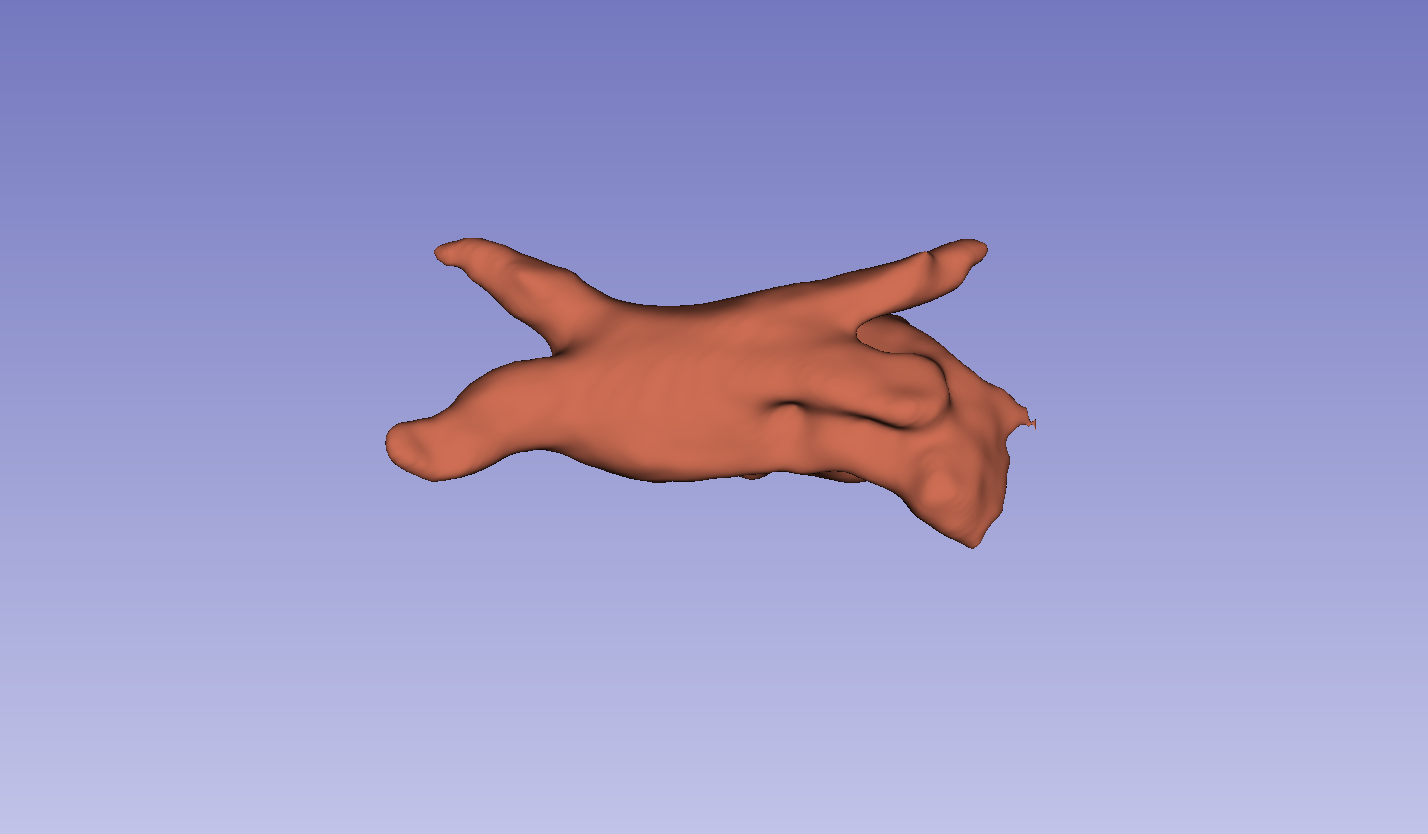


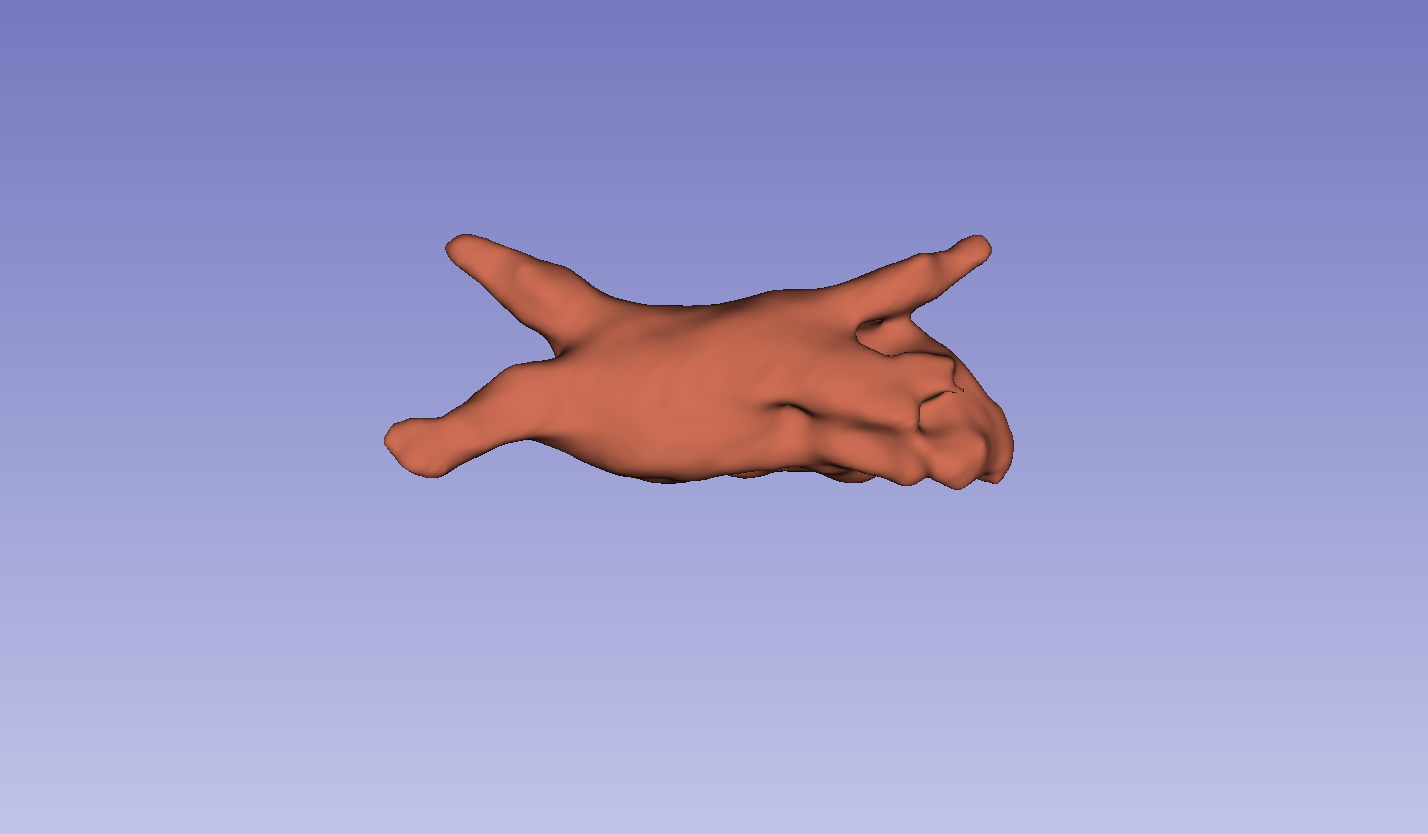

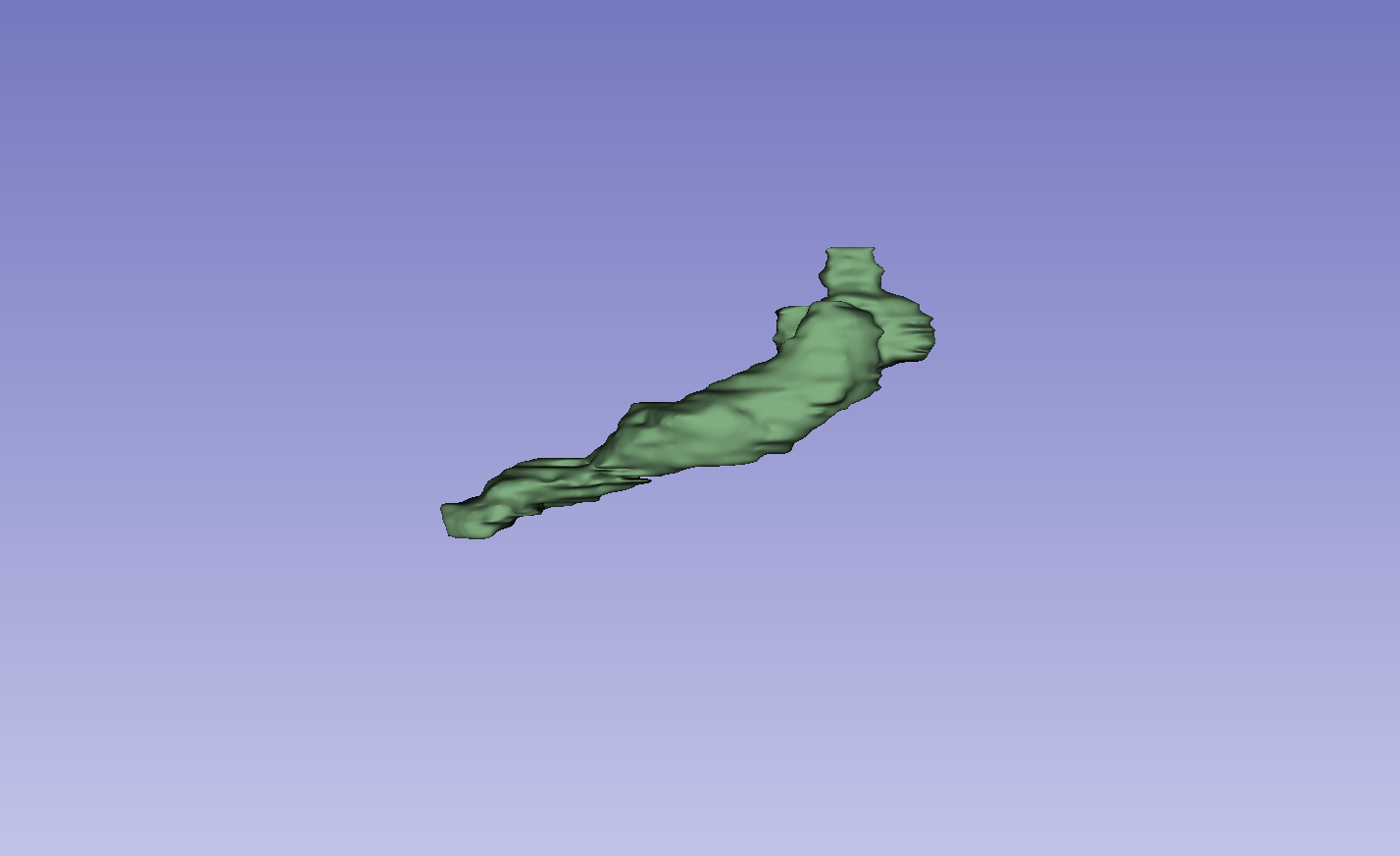
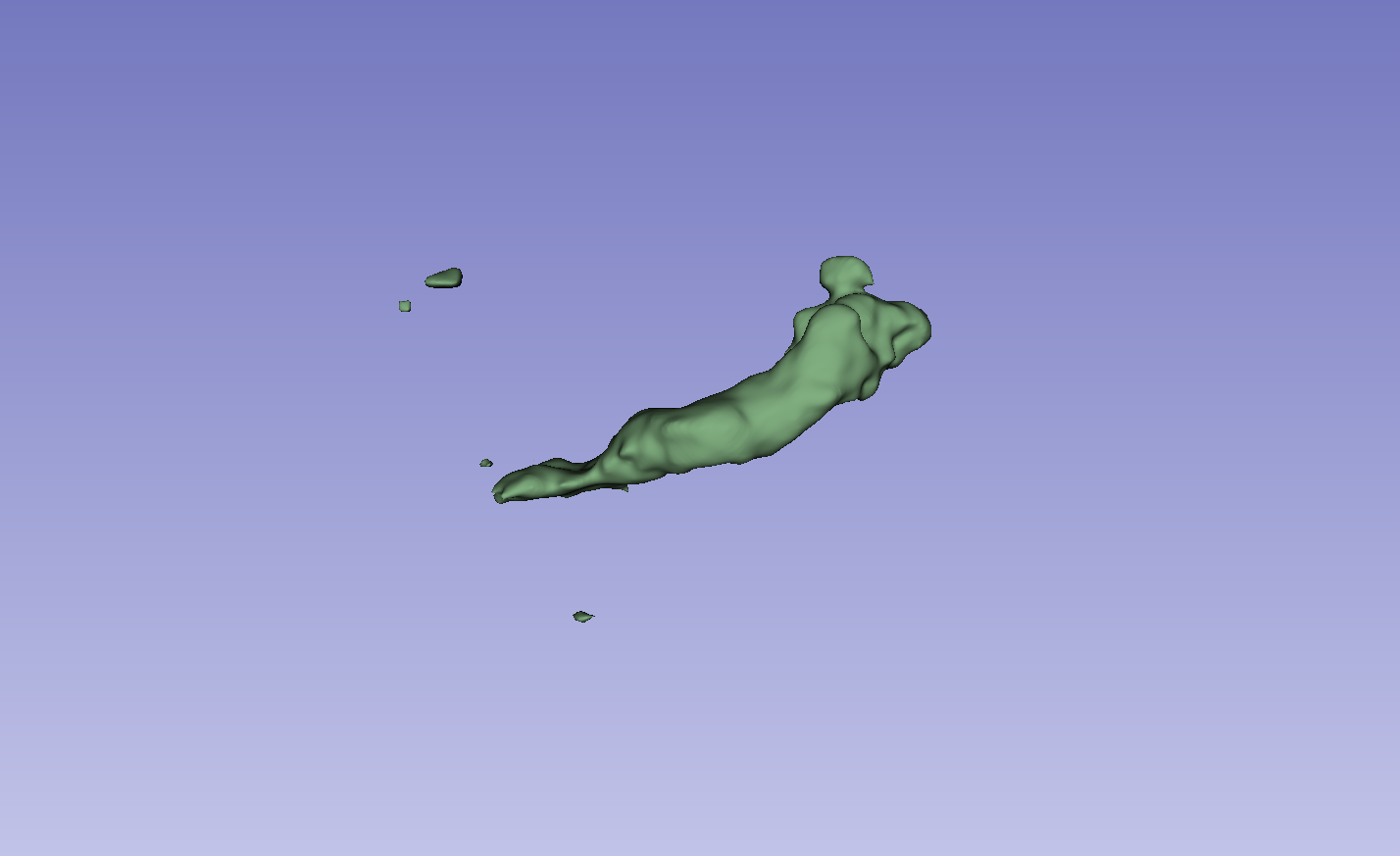
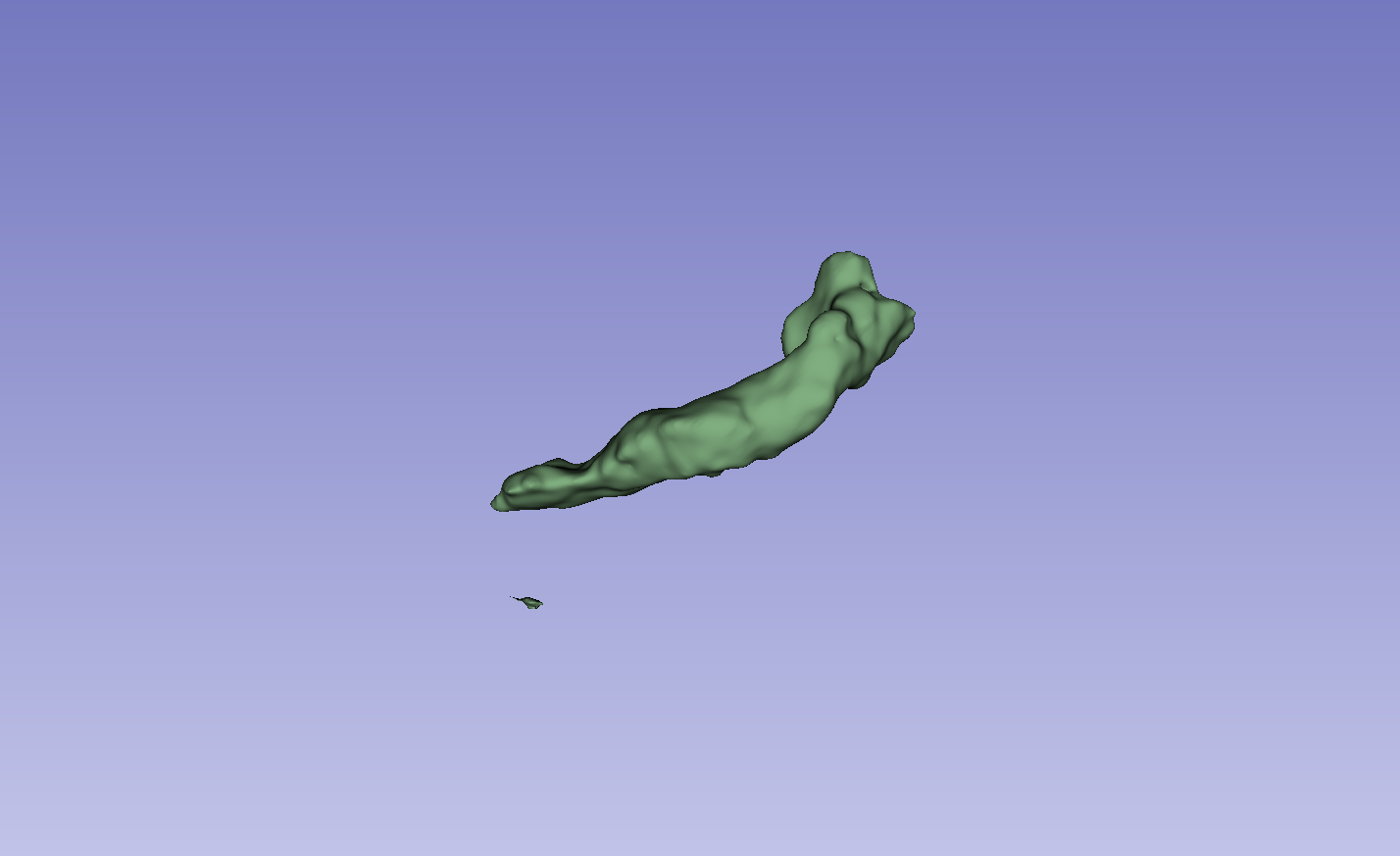
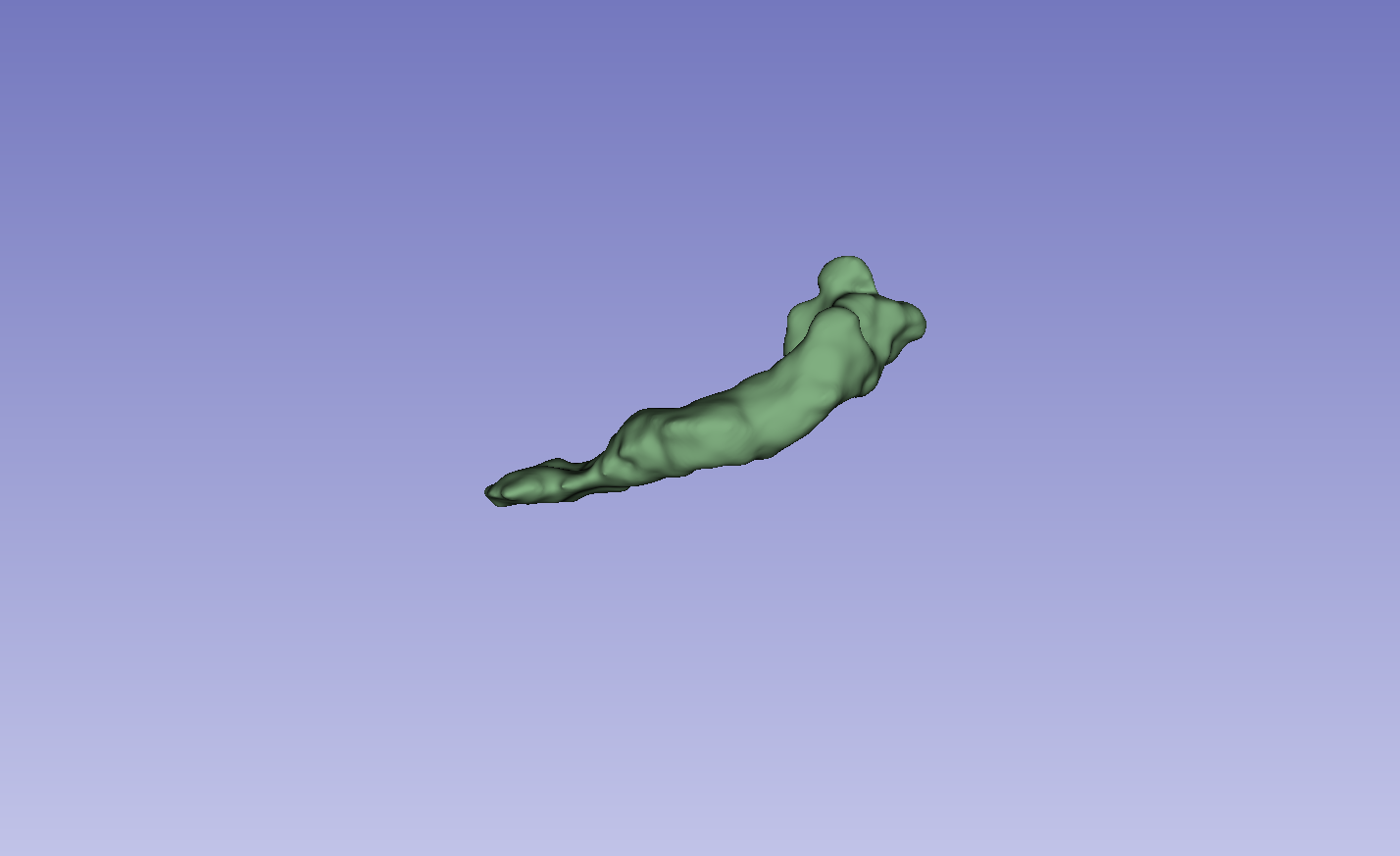
Figure 3: Visual comparison of segmentation results, demonstrating the method's ability to accurately segment the left atrium (LA) and pancreas compared to other methods.
The results on the LA dataset show improvements in all metrics, specifically DSC and Jaccard. Compared to MCF [wang2023mcf], the method exhibits an increase in DSC from 88.71 to 89.10 and Jaccard index from 80.41 to 81.62. The method also demonstrated strong performance on the Pancreas dataset. Visual results highlight higher overlap with ground truth labels and fewer false segmentations. Ablation studies on the LA dataset confirmed the impact of the regularization and contrastive loss components.
Conclusion
The dual-stream network with integrated contrastive learning and error correction mechanisms shows substantial improvements in semi-supervised semantic segmentation for medical imaging. The method's ability to effectively leverage both labeled and unlabeled data, while addressing the challenges of unreliable predictions, positions it as a valuable contribution to the field. The observed performance gains on the LA and NIH Pancreas datasets suggest the potential for broader applicability across various medical imaging tasks.









