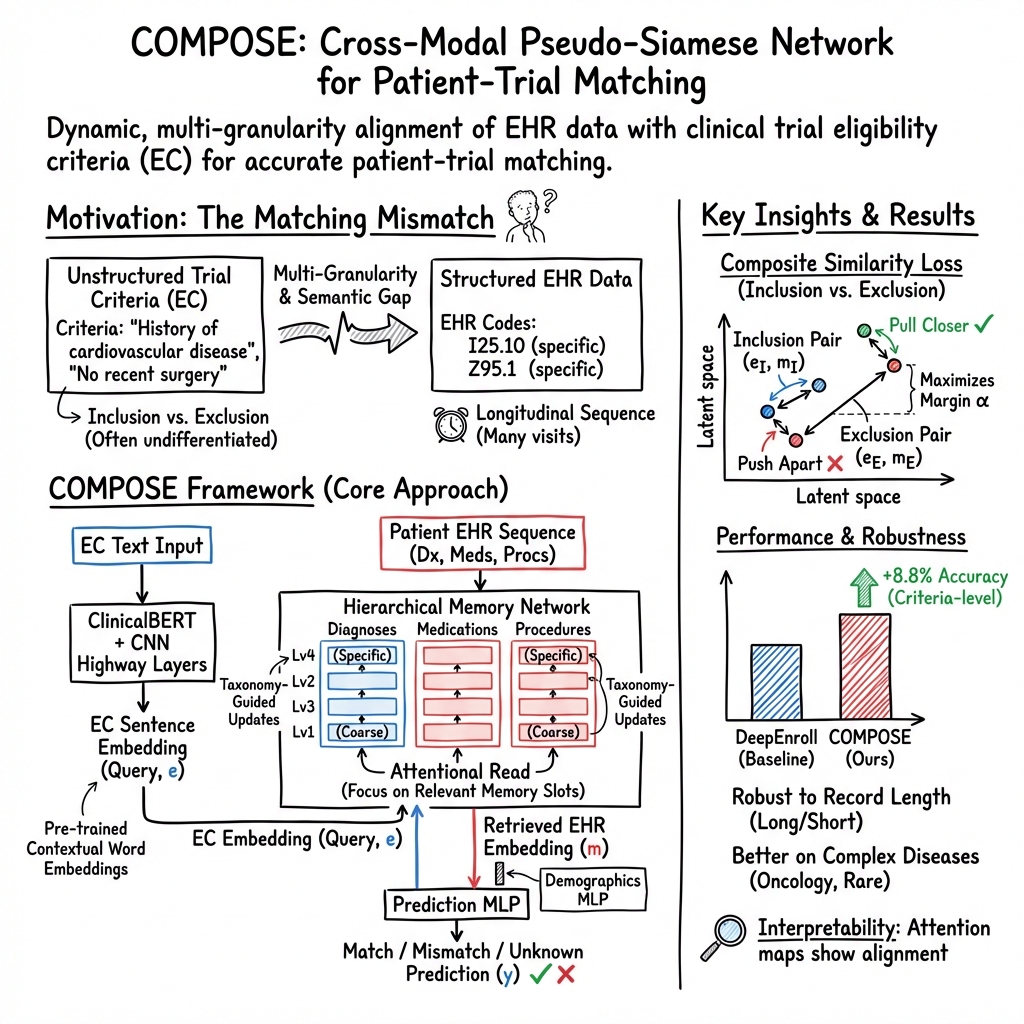
- The paper proposes COMPOSE, a cross-modal pseudo-siamese network that integrates heterogeneous medical data for precise patient-trial matching.
- It employs a dual-pathway architecture with a convolutional highway and a multi-granularity memory network to align EHR and trial eligibility criteria.
- Results show a 98% AUC and 83.7% accuracy, marking a 24.3% improvement over prior methods and reducing clinical trial recruitment inefficiencies.
COMPOSE: Cross-Modal Pseudo-Siamese Network for Patient Trial Matching
Introduction
The patient-trial matching problem deals with identifying suitable candidates for clinical trials using electronic health records (EHR) and trial eligibility criteria (ECs). The traditional patient recruitment process is plagued by inefficiencies and high costs, necessitating innovative computational methods to automate matching procedures. The COMPOSE model leverages the strengths of cross-modal learning and pseudo-siamese networks to advance this field, addressing key challenges such as the incorporation of heterogeneous medical concept granularity, many-to-many patient-trial relationships, and the explicit handling of inclusion and exclusion criteria in ECs.
Methods
COMPOSE employs a dual-pathway architecture: one pathway focuses on EC embedding using a convolutional highway network, while the other processes EHR data through a multi-granularity memory network. This novel approach integrates taxonomy-guided medical concept embedding to reconcile granularity discrepancies between detailed patient records and more general EC descriptions. Additionally, by utilizing attentive record alignment, COMPOSE dynamically matches patient records with trial criteria, effectively handling the distinct semantic roles of inclusion and exclusion criteria via a composite loss function. This design maximizes patient-record similarity with inclusion criteria and minimizes it with exclusion criteria.
Results
The COMPOSE model demonstrates superior performance over existing benchmarks in real-world datasets, achieving an area under the curve (AUC) of 98.0% for patient-criteria matching and an 83.7% accuracy for patient-trial matching. This marks a significant 24.3% improvement over the previous best methods. The results underscore COMPOSE's capability in effectively processing both structured and unstructured medical data and managing the complexities of clinical trial eligibility.
Implications
COMPOSE’s ability to handle diverse data modalities and its dynamic matching capabilities signify substantial progress towards automated, efficient patient-trial matching. The practical implications of these enhancements include reduced recruitment costs and timelines for clinical trials, potentially accelerating the drug development process. Theoretical implications further suggest that the dual pathway architecture and the incorporation of detailed medical taxonomies could benefit various tasks involving heterogeneous medical data.
Future Developments
Future research can extend COMPOSE by exploring its application across a broader spectrum of clinical trial phases and diverse medical conditions, including rare diseases. Enhancements could focus on refining the memory network for even finer-grained record alignment and exploring unsupervised or semi-supervised approaches to reduce labeled data dependency. Additionally, integrating real-time patient data updates could enhance COMPOSE’s dynamic matching capabilities, adapting criteria alignment as patient conditions evolve.
Conclusion
COMPOSE sets a new standard for patient-trial matching, leveraging cutting-edge cross-modal and pseudo-siamese network architectures to deliver substantial gains in matching accuracy and efficiency. Its success in the domain-specific challenges of clinical trials highlights a promising direction for computational methods in healthcare, potentially transforming patient recruitment processes and amplifying the efficiency of clinical research methodologies.