- The paper introduces CTP-LLM, a novel GPT-3.5-based framework for predicting clinical trial phase transitions using protocol documents.
- The methodology contrasts a fine-tuned GPT-3.5 model with a BERT+RF approach, achieving 67% overall and 75% Phase III prediction accuracy.
- The results imply that leveraging LLMs can streamline clinical trial evaluation, potentially reducing research time and optimizing resource allocation.
CTP-LLM: Clinical Trial Phase Transition Prediction Using LLMs
This essay provides a detailed examination of the "CTP-LLM: Clinical Trial Phase Transition Prediction Using LLMs," focusing on its methodologies, experimental results, and implications for the field of clinical trial prediction. The paper presents a novel approach that leverages LLMs, particularly a fine-tuned GPT-3.5 model, to predict clinical trial phase transitions, thereby offering insights into trial outcomes based on protocol documents.
Introduction and Methodology
The development of new medical treatments requires rigorous clinical trials, which are often characterized by high attrition rates. The research introduces a framework named CTP-LLM, the first LLM-based model specifically designed for Clinical Trial Outcome Prediction (CTOP) using trial protocol texts as input. This model eliminates the need for human-selected features, relying instead on the intrinsic ability of LLMs to extract relevant patterns from unstructured text data.
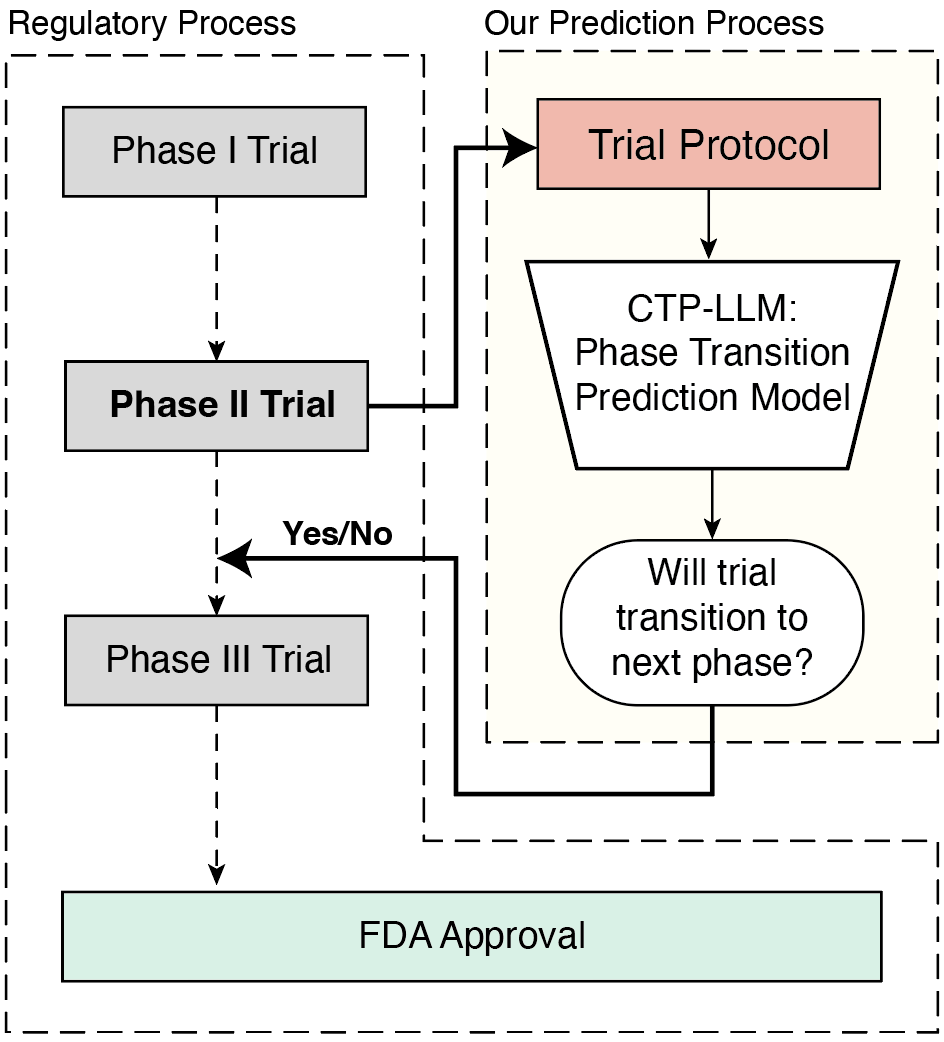
Figure 1: Overview of our framework, CTP-LLM, for clinical trial phase transition prediction. A trial protocol is a comprehensive document that outlines the plan for conducting the trial.
To implement this model, the authors built the PhaseTransition (PT) dataset, which labels clinical trials based on their progression through the regulatory phases. This dataset, coupled with the computational prowess of GPT-3.5, facilitates predicting whether a trial progresses to the next phase. The methodology involves two main models: CTP-LLM, a fine-tuned version of GPT-3.5, and BERT+RF, which combines clinical BERT with a Random Forest classifier.
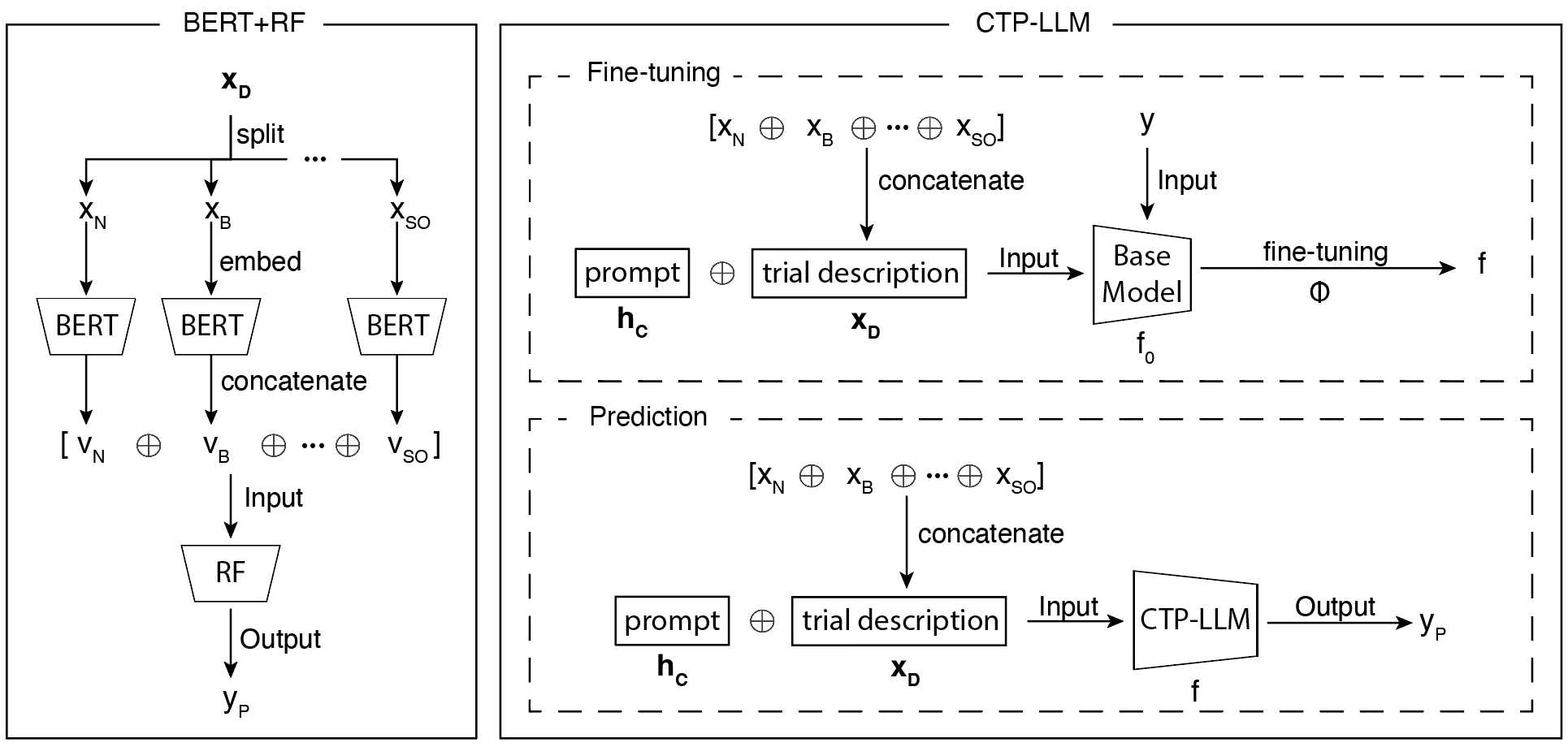
Figure 2: Overview of the two models. On the left is the BERT+RF approach, where the trial textual description is divided and individually embedded by clinical BERT before RF classification. On the right is the CTP-LLM approach with instruction fine-tuning of GPT-3.5.
Experimental Setup and Results
The paper reports that CTP-LLM achieves a 67% accuracy rate in predicting phase transitions across all phases, with a notable 75% accuracy rate for Phase III to approval transitions. BERT+RF, while advantageous for its lower computational cost, exhibited lower performance metrics compared to CTP-LLM. The dataset comprised 20,000 trial entries, and the experimental setup involved training, validation, and testing splits to ensure rigorous evaluation.
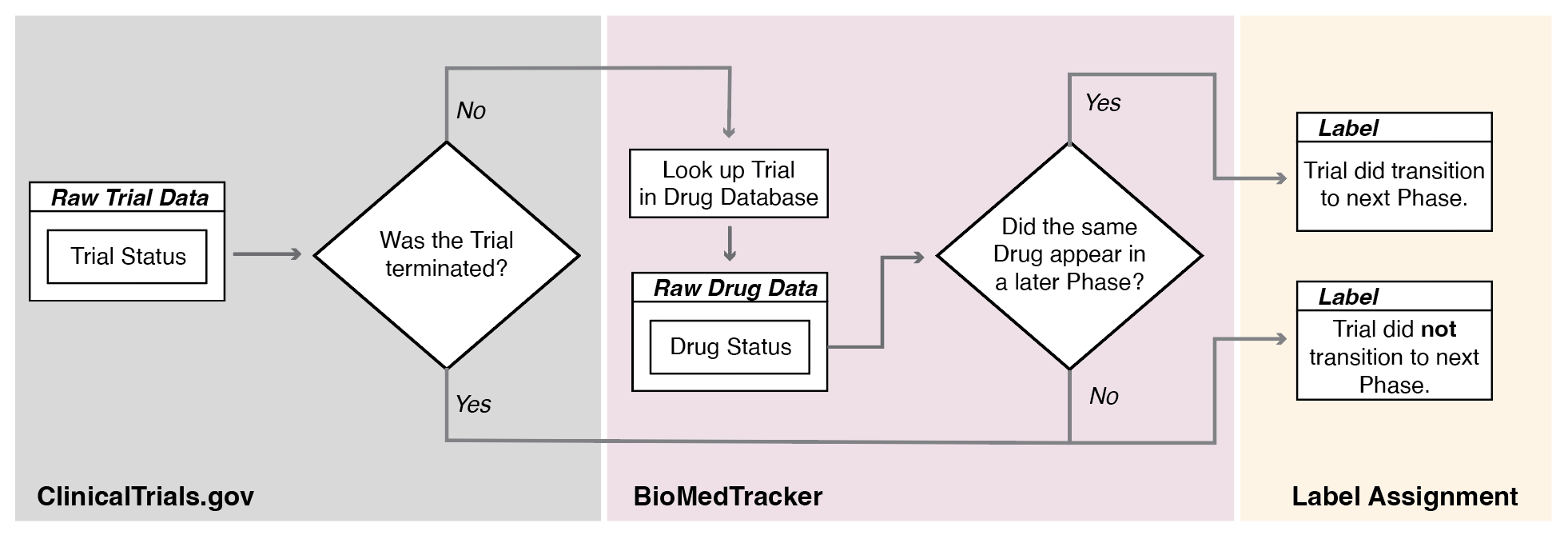
Figure 3: Overview of the labelling process, illustrating how trials are connected based on drug-indication ID for phase transition prediction.
The results demonstrate the potential application of LLMs in medical domains, offering a scalable solution for predicting trial outcomes. The fine-tuned model's ability to analyze complex protocol documents automatically and efficiently marks a step forward in clinical AI applications.
Implications and Future Directions
The research suggests that LLMs can significantly enhance the predictability of clinical trial outcomes, potentially reducing the time and cost associated with medical research. By predicting trial success early in the regulatory process, the CTP-LLM model could improve resource allocation and decision-making for trial designers.
The paper also contends with the limitations in trial data quality and underlines the necessity of comprehensive datasets for further advancements. Future research could explore the expansion of the current model to incorporate more diverse datasets, enhancing the model's robustness across different medical domains.
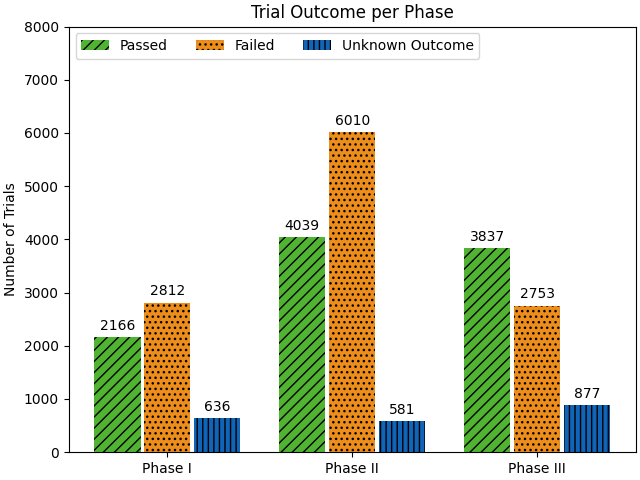
Figure 4: Distribution of Passed, Failed, and Unknown Outcome Trials, showing the highest attrition rate in Phase II.
In conclusion, the implementation of LLMs such as CTP-LLM in predicting clinical trial outcomes represents a promising advancement in leveraging AI for healthcare. With continued innovation and dataset refinement, LLMs could fundamentally transform the landscape of clinical trials, offering new efficiencies and insights into drug development processes.



