Towards an AI co-scientist
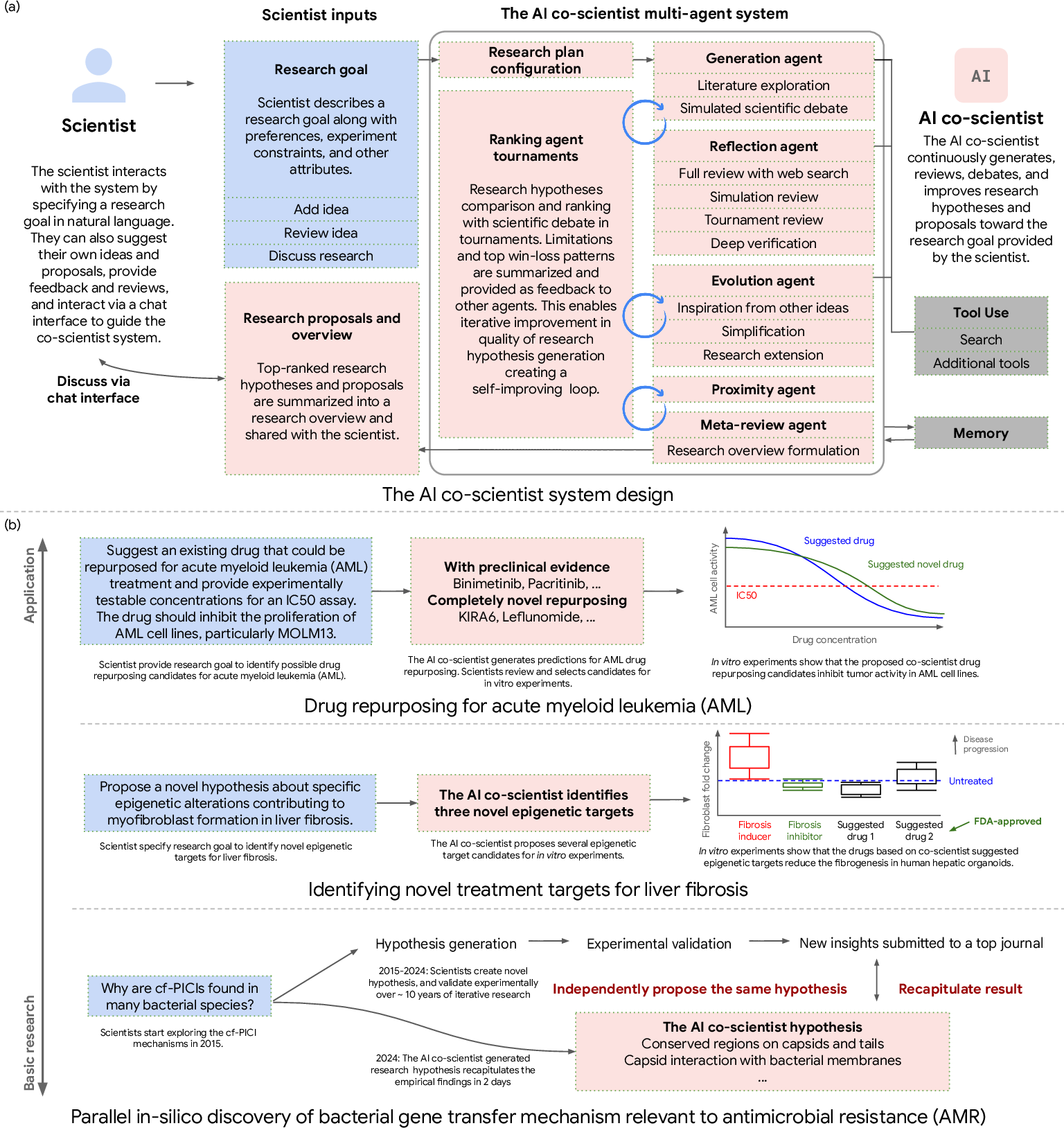
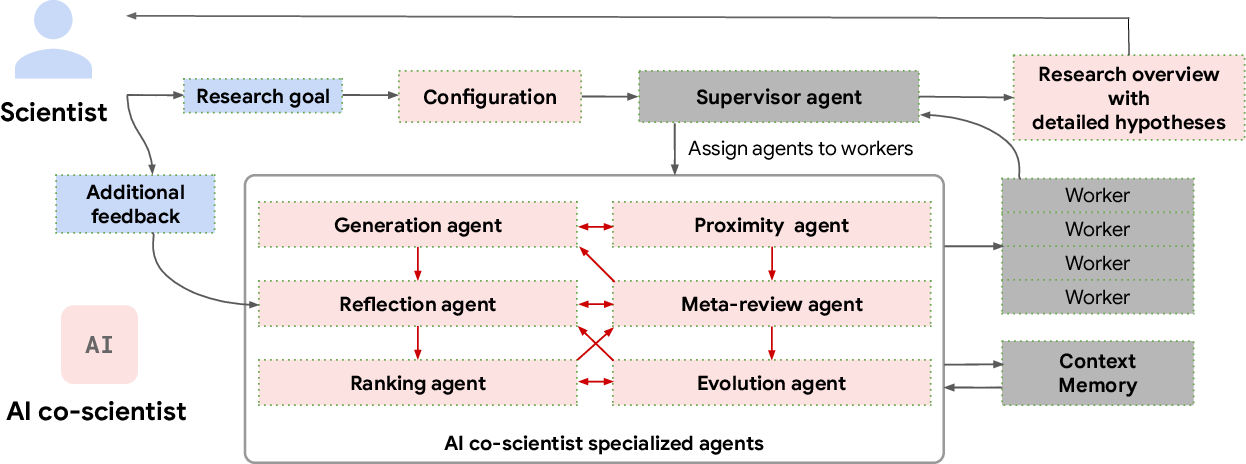
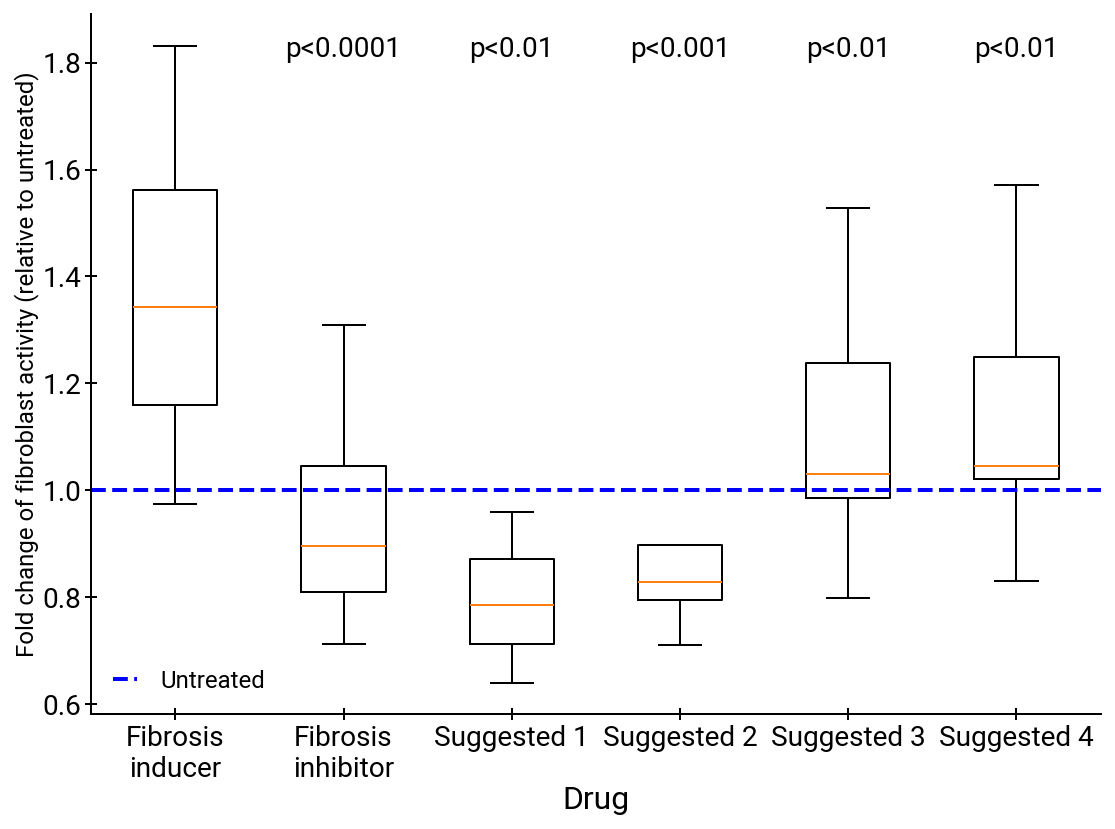
Abstract: Scientific discovery relies on scientists generating novel hypotheses that undergo rigorous experimental validation. To augment this process, we introduce an AI co-scientist, a multi-agent system built on Gemini 2.0. The AI co-scientist is intended to help uncover new, original knowledge and to formulate demonstrably novel research hypotheses and proposals, building upon prior evidence and aligned to scientist-provided research objectives and guidance. The system's design incorporates a generate, debate, and evolve approach to hypothesis generation, inspired by the scientific method and accelerated by scaling test-time compute. Key contributions include: (1) a multi-agent architecture with an asynchronous task execution framework for flexible compute scaling; (2) a tournament evolution process for self-improving hypotheses generation. Automated evaluations show continued benefits of test-time compute, improving hypothesis quality. While general purpose, we focus development and validation in three biomedical areas: drug repurposing, novel target discovery, and explaining mechanisms of bacterial evolution and anti-microbial resistance. For drug repurposing, the system proposes candidates with promising validation findings, including candidates for acute myeloid leukemia that show tumor inhibition in vitro at clinically applicable concentrations. For novel target discovery, the AI co-scientist proposed new epigenetic targets for liver fibrosis, validated by anti-fibrotic activity and liver cell regeneration in human hepatic organoids. Finally, the AI co-scientist recapitulated unpublished experimental results via a parallel in silico discovery of a novel gene transfer mechanism in bacterial evolution. These results, detailed in separate, co-timed reports, demonstrate the potential to augment biomedical and scientific discovery and usher an era of AI empowered scientists.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Glossary
- Acute myeloid leukemia (AML): A fast-growing cancer of the blood and bone marrow affecting myeloid lineage cells. "suggesting novel drug repurposing candidates for acute myeloid leukemia (AML) (upper panel)"
- Agentic behaviors: Autonomous, goal-directed actions by AI systems, including planning and tool use over long horizons. "agentic behaviors~\citep{wiesinger2024agents} such as the ability to use tools to solve complex tasks over long time horizons."
- Amyotrophic lateral sclerosis (ALS): A progressive neurodegenerative disease affecting motor neurons. "exploring the biological mechanisms of Amyotrophic Lateral Sclerosis (ALS)"
- Anti-fibrotic activity: Biological effects that reduce or prevent fibrosis (scar tissue formation). "validated by anti-fibrotic activity and liver cell regeneration in human hepatic organoids."
- Antimicrobial resistance (AMR): The ability of microbes to resist the effects of antimicrobials, undermining treatment. "antimicrobial resistance (AMR) - mechanisms developed by microbes to circumvent drug applications used to fight infections."
- Capsid-forming phage-inducible chromosomal islands (cf-PICIs): Mobile genetic elements that hijack phage capsid systems to package and transfer their genomes between bacteria. "The system was instructed to hypothesize how capsid-forming phage-inducible chromosomal islands (cf-PICIs) exist across multiple bacterial species."
- Chain-of-thought: Step-by-step reasoning traces produced by a model to solve complex problems. "leveraging reinforcement learning to refine the model's ``chain-of-thought'' and enhance complex reasoning abilities over longer horizons."
- Cell line: A population of cells grown in vitro that can be maintained over time for experiments. "across multiple AML cell lines."
- Conjugation: A mechanism of horizontal gene transfer in bacteria via direct cell-to-cell contact. "the molecular mechanisms of gene transfer (conjugation, transduction, and transformation)"
- Deep verification review: A rigorous decomposition and evaluation of a hypothesis into assumptions and sub-assumptions to identify failures. "The Reflection agent also conducts a deep verification review, decomposing the hypothesis into constituent assumptions."
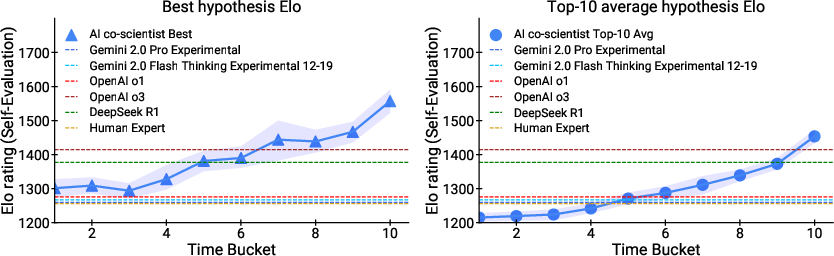
- Elo-based tournament: A ranking framework using the Elo rating system to compare and prioritize competing hypotheses. "The Ranking agent employs and orchestrates an Elo-based tournament~\cite{elo1978rating} to assess and prioritize the generated hypotheses at any given time."
- Epigenetic targets: Genes or regulatory elements modifiable without changing DNA sequence, implicated for therapeutic intervention. "new epigenetic targets for liver fibrosis, validated by anti-fibrotic activity and liver cell regeneration in human hepatic organoids."
- Graph convolutional networks: Neural networks that operate on graph-structured data, aggregating information from neighbors. "knowledge graphs with graph convolutional networks"
- Hepatic organoids: Three-dimensional, lab-grown mini-organs derived from liver cells that model liver function. "liver cell regeneration in human hepatic organoids."
- Host range: The spectrum of hosts a pathogen or genetic element can infect or operate within. "cf-PICIs interact with diverse phage tails to expand their host range."
- In silico: Performed via computer simulation or computational methods rather than in the lab or living organisms. "a parallel in silico discovery of a novel gene transfer mechanism in bacterial evolution."
- In vitro: Experiments conducted outside a living organism, typically in test tubes or culture dishes. "that show tumor inhibition in vitro at clinically applicable concentrations."
- In vivo: Experiments conducted within living organisms. "in vitro and in vivo experimentation."
- Inductive biases: Assumptions built into models or algorithms that guide learning and reasoning. "using inductive biases derived from the scientific method to design a multi-agent framework for scientific reasoning"
- Knowledge graph: A structured representation of entities and their relationships used for reasoning and inference. "remains dependent on the underlying knowledge graph's quality and lacks sufficient scalability and explainability."
- Monte Carlo Tree Search (MCTS): A heuristic search algorithm using randomized simulations to explore decision trees. "used Monte Carlo Tree Search (MCTS) to explore game states and strategically select moves"
- Murine: Pertaining to mice; commonly used in biomedical research contexts. "identify causative murine genetic factors for traits such as diabetes, cataracts, and hearing loss"
- Nuclear Pore Complex (NPC): Large protein assemblies in the nuclear envelope that regulate transport between nucleus and cytoplasm. "phosphorylation of the Nuclear Pore Complex (NPC)"
- Phage tails: Tail structures of bacteriophages used for host recognition and DNA injection. "cf-PICIs interact with diverse phage tails to expand their host range."
- Phosphorylation: The addition of a phosphate group to a molecule, often regulating protein function. "phosphorylation of the Nuclear Pore Complex (NPC)"
- Principal investigator (PI): The lead researcher responsible for the design and conduct of a project. "a ``principal investigator'' LLM guiding a team of specialized LLM agents"
- Proximity graph: A graph connecting items based on similarity or distance to enable clustering and exploration. "This agent asynchronously computes a proximity graph for generated hypotheses, enabling clustering of similar ideas, de-duplication, and efficient exploration of the hypothesis landscape."
- Self-play: A training or reasoning strategy where an agent competes or debates with versions of itself to improve. "a self-play based scientific debate step for generating novel research hypotheses"
- Test-time compute paradigm: Allocating additional computation during inference to enable deeper reasoning and better performance. "One promising direction in this pursuit is the test-time compute paradigm."
- Transduction: Transfer of genetic material between bacteria mediated by bacteriophages. "the molecular mechanisms of gene transfer (conjugation, transduction, and transformation)"
- Transformation: Uptake of free DNA from the environment by bacteria, leading to genetic change. "the molecular mechanisms of gene transfer (conjugation, transduction, and transformation)"
- Wet lab: Laboratory work involving physical experiments with biological or chemical materials. "validated in wet-lab settings"
- Zero-shot: Generalization to novel tasks or classes without task-specific training examples. "addresses ``zero-shot'' repurposing for novel diseases"
Collections
Sign up for free to add this paper to one or more collections.