- The paper proposes a novel DISTL framework that leverages self-supervised learning and a Vision Transformer to achieve 98.14% accuracy in TB screening.
- It utilizes U-Net segmentation on chest x-ray images to focus on lung regions, thereby enhancing diagnostic precision and recall in settings with scarce labeled data.
- Explainability analysis with visual heatmaps confirms that the model accurately identifies key TB indicators, aiding rapid and reliable clinical decision-making.
Empowering Tuberculosis Screening with Explainable Self-Supervised Deep Neural Networks
Introduction
The paper "Empowering Tuberculosis Screening with Explainable Self-Supervised Deep Neural Networks" (2406.13750) addresses the persistent global health challenge posed by tuberculosis (TB), particularly in resource-constrained settings. TB remains an acute issue, primarily affecting economically disadvantaged regions, despite being a curable and manageable disease. The diagnostic process for TB heavily relies on chest x-ray (CXR) imaging, which requires skilled radiologists whose availability is limited in many areas. This situation emphasizes the need for artificial intelligence-powered solutions to assist clinicians in the rapid screening and diagnosis of TB.
The authors present a novel AI framework based on explainable self-supervised learning to improve TB screening efficacy. The network achieves remarkable performance metrics, with an overall accuracy of 98.14%, a recall rate of 95.72%, and a precision rate of 99.44% in detecting TB cases. These results underscore the network's capability to capture clinically significant features in CXR images.
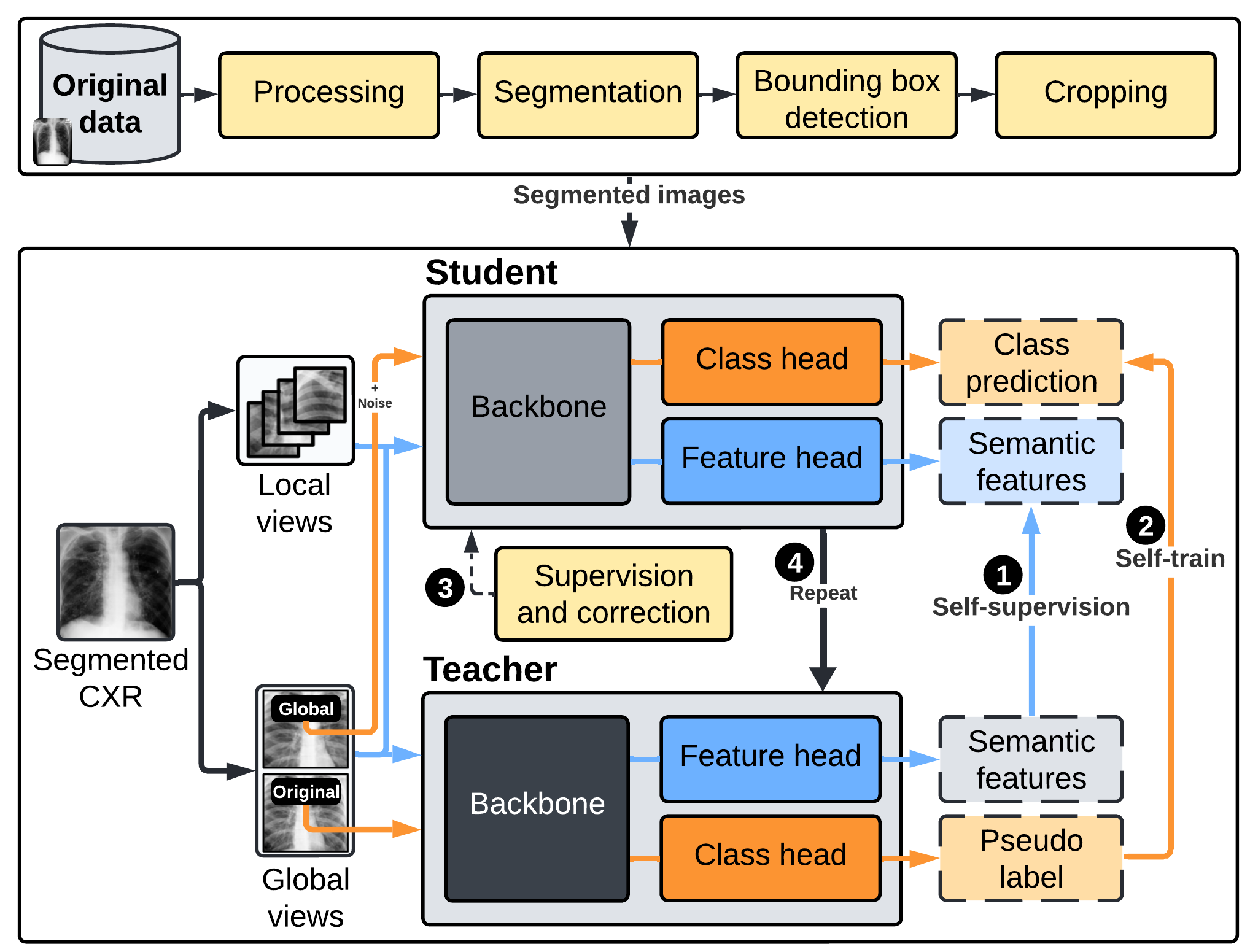
Figure 1: The high-level conceptual flow of the framework that encompasses three primary components for segmentation, self-supervision, and self-training.
Methodology
Data Collection and Preparation
The research utilized a diverse dataset comprising chest x-ray images collected from four sources: Montgomery, Shenzhen, Belarus, and Rahman et al. datasets. The final dataset included 1,923 tuberculosis-positive images and 2,984 normal images, following rigorous preprocessing and quality assurance steps. The segmentation of lung regions was achieved using the U-Net model to focus subsequent analyses exclusively on relevant regions, increasing the segmentation accuracy and excluding irrelevant artifacts.
The dataset was divided into labeled and unlabeled subsets, facilitating both self-supervised and supervised learning phases.
Network Architecture
The AI framework employs a distillation approach for self-supervised and self-trained learning known as DISTL. DISTL mimics radiologists' learning by integrating self-supervisory fine-tuned techniques to identify patterns in medical images without large labeled datasets. The system architecture utilizes a Vision Transformer (ViT) backbone with innovate features such as multilayer perceptrons and self-supervision via Distillation with No Labels (DINO). The framework is designed with a focus on capturing holistic image cues through self-attention mechanisms, a superior alternative to convolution-based models.

Figure 1: The high-level conceptual flow of the framework that encompasses three primary components for segmentation, self-supervision, and self-training.
Training and Evaluation
The network undergoes multiple training iterations using a combination of labeled and unlabeled data, employing advanced data augmentations to enhance robustness. Performance metrics like recall, precision, and accuracy were evaluated alongside convolutional network baselines, demonstrating superior outcomes with the novel approach. The AI model development environment utilized PyTorch, Pillow, OpenCV-Python, and Scikit-learn, powered by an NVIDIA RTX GPU for efficient computation.
The explainability of the model was assessed, verifying that relevant radiographic indicators for TB were accurately identified, highlighting the model's potential in supporting clinicians with evidence-backed AI screening tools.
Results
The proposed system demonstrated distinguished performance compared to existing convolutional neural network models such as vanilla CNN, VGG16, ResNet18, and ResNet50. Key metrics such as accuracy, precision, and recall showed marked improvement over these benchmarks. Precision for detecting TB was recorded at 99.44%, and normal cases were identified with a precision of 97.37%, which outperformed traditional models across all metrics.
Explainability Analysis
The attention-driven framework was validated through explainability analysis, illustrating reliance on medically pertinent lung regions and rejecting erroneous or non-relevant indicators. Visual heatmaps further supported these findings. Critical TB indicators such as infiltrates, cavitation, and consolidations were accurately localized, consistent with clinical diagnostics.
(Figure 2)
Figure 2: Sample tuberculosis-positive cases with identified critical regions.
Conclusion
The paper demonstrates a viable approach to enhancing TB screening using explainable, self-supervised deep learning models. The proposed DISTL-based methodology combines self-supervision with self-training to achieve high diagnostic performance with limited labeled data. Future research should aim to broaden data diversity and include clinical validation across varied populations.
In the context of global health, this research underscores the promise of leveraging AI to address resource scarcity, offering clinicians rapid diagnostic assistance in TB endemic areas. Continued work will involve validation with certified radiologists and expansion to larger datasets, enhancing the universal applicability and reliability of AI-driven healthcare solutions.
