GFlowNets for AI-Driven Scientific Discovery
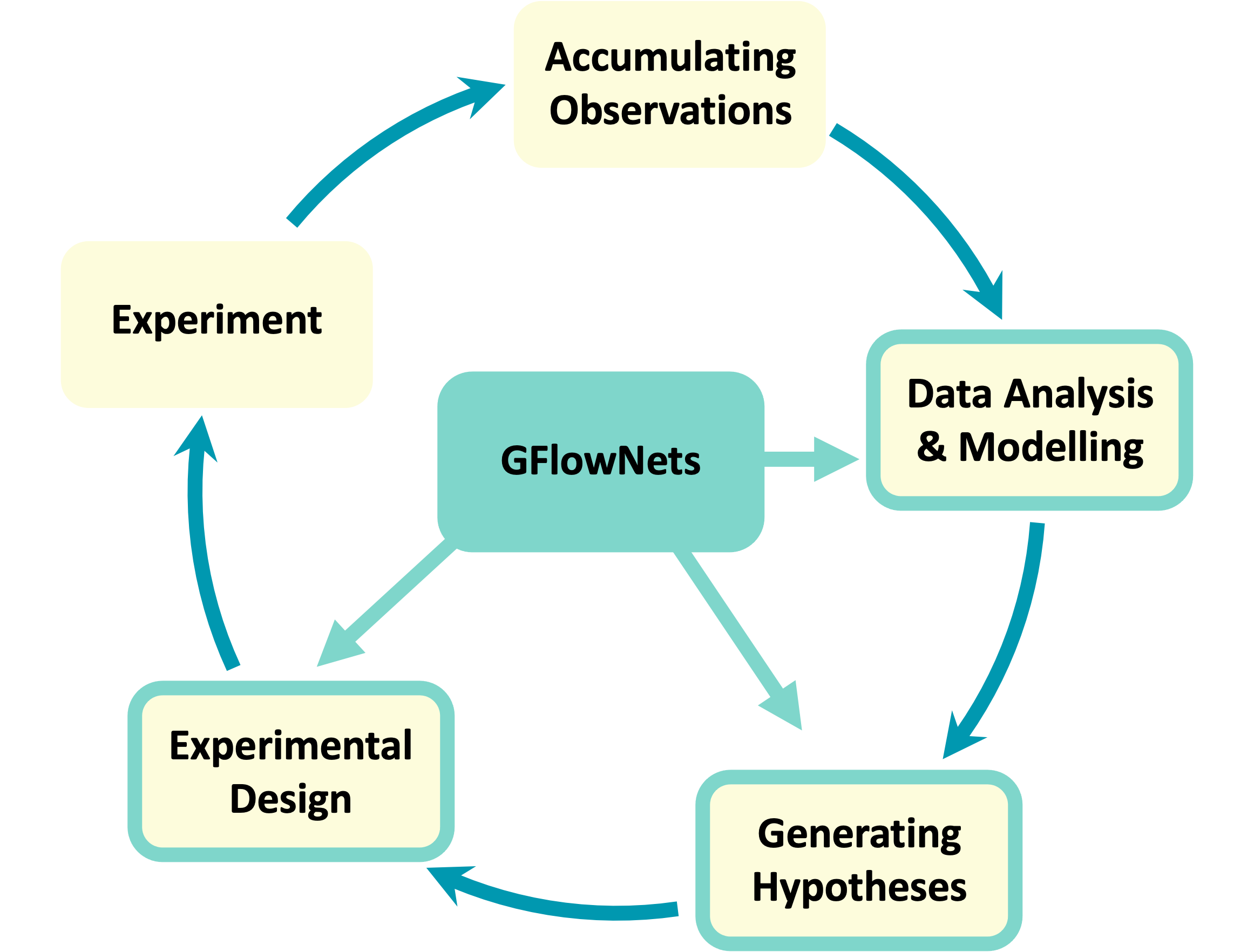
Abstract: Tackling the most pressing problems for humanity, such as the climate crisis and the threat of global pandemics, requires accelerating the pace of scientific discovery. While science has traditionally relied on trial and error and even serendipity to a large extent, the last few decades have seen a surge of data-driven scientific discoveries. However, in order to truly leverage large-scale data sets and high-throughput experimental setups, machine learning methods will need to be further improved and better integrated in the scientific discovery pipeline. A key challenge for current machine learning methods in this context is the efficient exploration of very large search spaces, which requires techniques for estimating reducible (epistemic) uncertainty and generating sets of diverse and informative experiments to perform. This motivated a new probabilistic machine learning framework called GFlowNets, which can be applied in the modeling, hypotheses generation and experimental design stages of the experimental science loop. GFlowNets learn to sample from a distribution given indirectly by a reward function corresponding to an unnormalized probability, which enables sampling diverse, high-reward candidates. GFlowNets can also be used to form efficient and amortized Bayesian posterior estimators for causal models conditioned on the already acquired experimental data. Having such posterior models can then provide estimators of epistemic uncertainty and information gain that can drive an experimental design policy. Altogether, here we will argue that GFlowNets can become a valuable tool for AI-driven scientific discovery, especially in scenarios of very large candidate spaces where we have access to cheap but inaccurate measurements or to expensive but accurate measurements. This is a common setting in the context of drug and material discovery, which we use as examples throughout the paper.
- Cross-Chapter Paper 5: Mountains, pages 2273–2318. Cambridge University Press, Cambridge, UK and New York, USA, 2022. ISBN 9781009325844. doi: 10.1017/9781009325844.022.2273.
- Thomas A Ban. The role of serendipity in drug discovery. Dialogues in clinical neuroscience, 2022.
- The art and practice of structure-based drug design: A molecular modeling perspective. Medicinal Research Reviews, 16(1):3–50, 1996. doi: https://doi.org/10.1002/(SICI)1098-1128(199601)16:1¡3::AID-MED1¿3.0.CO;2-6. URL https://onlinelibrary.wiley.com/doi/abs/10.1002/%28SICI%291098-1128%28199601%2916%3A1%3C3%3A%3AAID-MED1%3E3.0.CO%3B2-6.
- Self-driving laboratory for accelerated discovery of thin-film materials. Science Advances, 6(20):eaaz8867, 2020.
- The fourth paradigm: data-intensive scientific discovery, volume 1. Microsoft research Redmond, WA, 2009.
- Perspective: Materials informatics and big data: Realization of the “fourth paradigm” of science in materials science. Apl Materials, 4(5):053208, 2016.
- 3d infomax improves gnns for molecular property prediction. In International Conference on Machine Learning, pages 20479–20502. PMLR, 2022.
- A review of modern computational algorithms for bayesian optimal design. International Statistical Review, 84(1):128–154, 2016.
- Model-based reinforcement learning for biological sequence design. In International conference on learning representations, 2019.
- Deep learning for bayesian optimization of scientific problems with high-dimensional structure. Transactions of Machine Learning Research, 2022.
- Flow network based generative models for non-iterative diverse candidate generation. In A. Beygelzimer, Y. Dauphin, P. Liang, and J. Wortman Vaughan, editors, Advances in Neural Information Processing Systems, 2021a. URL https://openreview.net/forum?id=Arn2E4IRjEB.
- Gflownet foundations, 2021b.
- Mastering the game of go with deep neural networks and tree search. Nature, 529(7587):484–489, 2016.
- Highly accurate protein structure prediction with alphafold. Nature, 596(7873):583–589, 2021.
- Aleatory or epistemic? does it matter? Structural Safety, 31(2):105–112, 2009. ISSN 0167-4730. doi: https://doi.org/10.1016/j.strusafe.2008.06.020. Risk Acceptance and Risk Communication.
- Discovery of novel li sse and anode coatings using interpretable machine learning and high-throughput multi-property screening. Scientific Reports, 11(1):1–14, 2021.
- Toward causal representation learning. Proceedings of the IEEE, 109(5):612–634, 2021.
- An introduction to mcmc for machine learning. Machine learning, 50(1):5–43, 2003.
- Nonphysical sampling distributions in monte carlo free-energy estimation: Umbrella sampling. Journal of Computational Physics, 23(2):187–199, 1977.
- Parallel tempering: Theory, applications, and new perspectives. Physical Chemistry Chemical Physics, 7(23):3910–3916, 2005.
- Hybrid monte carlo on hilbert spaces. Stochastic Processes and their Applications, 121(10):2201–2230, 2011.
- Variational inference: A review for statisticians. Journal of the American statistical Association, 112(518):859–877, 2017.
- Tom Minka et al. Divergence measures and message passing. Technical report, Technical report, Microsoft Research, 2005.
- Two problems with variational expectation maximisation for time series models, page 104–124. Cambridge University Press, 2011. doi: 10.1017/CBO9780511984679.006.
- Gflownets and variational inference. In International Conference on Learning Representations (ICLR), 2023.
- Valerii Vadimovich Fedorov. Theory of optimal experiments. Elsevier, 1972.
- Optimum experimental designs, volume 5. Clarendon Press, 1992.
- Burr Settles. Active learning. Synthesis lectures on artificial intelligence and machine learning, 6(1):1–114, 2012.
- Bayesian experimental design: A review. Statistical Science, pages 273–304, 1995.
- Dennis V Lindley. On a measure of the information provided by an experiment. The Annals of Mathematical Statistics, 27(4):986–1005, 1956.
- A tutorial on adaptive design optimization. Journal of mathematical psychology, 57(3-4):53–67, 2013.
- On nesting monte carlo estimators. In International Conference on Machine Learning, pages 4267–4276. PMLR, 2018.
- Variational bayesian optimal experimental design. Advances in Neural Information Processing Systems, 32, 2019.
- Efficient bayesian experimental design for implicit models. In The 22nd International Conference on Artificial Intelligence and Statistics, pages 476–485. PMLR, 2019.
- Bayesian experimental design for implicit models by mutual information neural estimation. In International Conference on Machine Learning, pages 5316–5326. PMLR, 2020.
- Information gain from isotopic contrast variation in neutron reflectometry on protein–membrane complex structures. Journal of applied crystallography, 53(3):800–810, 2020.
- Bayesian design of experiments for intractable likelihood models using coupled auxiliary models and multivariate emulation. Bayesian Analysis, 15(1):103–131, 2020.
- Sequential monte carlo for bayesian sequentially designed experiments for discrete data. Computational Statistics & Data Analysis, 57(1):320–335, 2013.
- A unified stochastic gradient approach to designing bayesian-optimal experiments. In International Conference on Artificial Intelligence and Statistics, pages 2959–2969. PMLR, 2020.
- Deep adaptive design: Amortizing sequential bayesian experimental design. In International Conference on Machine Learning, pages 3384–3395. PMLR, 2021.
- Implicit deep adaptive design: policy-based experimental design without likelihoods. Advances in Neural Information Processing Systems, 34:25785–25798, 2021.
- Optimizing sequential experimental design with deep reinforcement learning. In International Conference on Machine Learning, pages 2107–2128. PMLR, 2022.
- Introduction to global optimization. Springer Science & Business Media, 2000.
- The application of bayesian methods for seeking the extremum. Towards global optimization, 2(117-129):2, 1978.
- Efficient global optimization of expensive black-box functions. Journal of Global optimization, 13(4):455–492, 1998.
- Roman Garnett. Bayesian Optimization. Cambridge University Press, 2022. in preparation.
- Bayesian optimization for synthetic gene design. arXiv preprint arXiv:1505.01627, 2015.
- Constrained bayesian optimization for automatic chemical design using variational autoencoders. Chemical science, 11(2):577–586, 2020.
- Gaussian process molecule property prediction with flowmo. arXiv preprint arXiv:2010.01118, 2020.
- Bayesian reaction optimization as a tool for chemical synthesis. Nature, 590(7844):89–96, 2021.
- Gaussian Processes for Machine Learning. Adaptive Computation and Machine Learning series. MIT Press, 2005. ISBN 9780262182539. URL https://books.google.ca/books?id=GhoSngEACAAJ.
- Botorch: a framework for efficient monte-carlo bayesian optimization. Advances in neural information processing systems, 33:21524–21538, 2020.
- Bayesian approach to global optimization and application to multiobjective and constrained problems. Journal of optimization theory and applications, 70(1):157–172, 1991.
- Gaussian process optimization in the bandit setting: no regret and experimental design. In Proceedings of the 27th International Conference on International Conference on Machine Learning, pages 1015–1022, 2010.
- William R Thompson. On the likelihood that one unknown probability exceeds another in view of the evidence of two samples. Biometrika, 25(3-4):285–294, 1933.
- Scalable thompson sampling using sparse gaussian process models. Advances in Neural Information Processing Systems, 34:5631–5643, 2021.
- Entropy search for information-efficient global optimization. Journal of Machine Learning Research, 13(6), 2012.
- Predictive entropy search for efficient global optimization of black-box functions. Advances in neural information processing systems, 27, 2014.
- Output-space predictive entropy search for flexible global optimization. In NIPS workshop on Bayesian Optimization, pages 1–5, 2015.
- Zi Wang and Stefanie Jegelka. Max-value entropy search for efficient bayesian optimization. In International Conference on Machine Learning, pages 3627–3635. PMLR, 2017.
- Gibbon: General-purpose information-based bayesian optimisation. Journal of Machine Learning Research, 22(235):1–49, 2021.
- Batch bayesian optimization via local penalization. In Artificial intelligence and statistics, pages 648–657. PMLR, 2016.
- Parallelised bayesian optimisation via thompson sampling. In International Conference on Artificial Intelligence and Statistics, pages 133–142. PMLR, 2018.
- Noisy expected improvement and on-line computation time allocation for the optimization of simulators with tunable fidelity, 2010.
- Multi-fidelity bayesian optimisation with continuous approximations. In International Conference on Machine Learning, pages 1799–1808. PMLR, 2017.
- Multi-fidelity bayesian optimization with max-value entropy search and its parallelization. In International Conference on Machine Learning, pages 9334–9345. PMLR, 2020.
- Max-value entropy search for multi-objective bayesian optimization. Advances in Neural Information Processing Systems, 32, 2019.
- Multi-objective bayesian optimization over high-dimensional search spaces. In Uncertainty in Artificial Intelligence, pages 507–517. PMLR, 2022.
- Hypothesis learning in automated experiment: application to combinatorial materials libraries. Advanced Materials, 34(20):2201345, 2022.
- On-the-fly autonomous control of neutron diffraction via physics-informed bayesian active learning. Applied Physics Reviews, 9(2):021408, 2022.
- Boss: Bayesian optimization over string spaces. Advances in neural information processing systems, 33:15476–15486, 2020.
- Amortized bayesian optimization over discrete spaces. In Conference on Uncertainty in Artificial Intelligence, pages 769–778. PMLR, 2020.
- Judea Pearl. Probabilistic Reasoning in Intelligent Systems. Morgan Kaufmann, 1988.
- Causation, Prediction, and Search. MIT press, 2000.
- David Maxwell Chickering. Optimal Structure Identification With Greedy Search. Journal of Machine Learning Research, 2002.
- DAGs with NO TEARS: Continuous Optimization for Structure Learning. In Advances in Neural Information Processing Systems, 2018.
- Learning neural causal models from unknown interventions. arXiv preprint, 2019.
- Differentiable Causal Discovery from Interventional Data. Advances in Neural Information Processing Systems, 2020.
- Bayesian Graphical Models for Discrete Data. International Statistical Review, 1995.
- Being Bayesian About Network Structure. A Bayesian Approach to Structure Discovery in Bayesian Networks. Machine Learning, 2003.
- Improving Markov chain Monte Carlo model search for data mining. Machine learning, 2003.
- Structure Discovery in Bayesian Networks by Sampling Partial Orders. The Journal of Machine Learning Research, 2016.
- Towards Scalable Bayesian Learning of Causal DAGs. Advances in Neural Information Processing Systems, 2020.
- Data Analysis with Bayesian Networks: A Bootstrap Approach. Proceedings of the Fifteenth conference on Uncertainty in Artificial Intelligence, 1999.
- Abcd-strategy: Budgeted experimental design for targeted causal structure discovery. In The 22nd International Conference on Artificial Intelligence and Statistics. PMLR, 2019.
- Variational Causal Networks: Approximate Bayesian Inference over Causal Structures. arXiv preprint, 2021.
- BCD Nets: Scalable Variational Approaches for Bayesian Causal Discovery. Advances in Neural Information Processing Systems, 2021.
- Tractable Uncertainty for Structure Learning. International Conference on Machine Learning, 2022.
- DiBS: Differentiable Bayesian Structure Learning. Advances in Neural Information Processing Systems, 2021.
- Amortized Inference for Causal Structure Learning. Advances in Neural Information Processing Systems, 2022a.
- W BUNTINE. Theory refinement on bayesian networks. In Proc. 7th Conf. Uncertainty in Artificial Intelligence, 1991, pages 52–60, 1991.
- Active learning for structure in Bayesian networks. International Joint Conference on Artificial Intelligence, 2001.
- Kevin Murphy. Active Learning of Causal Bayes Net Structure, 2001.
- Learning Neural Causal Models with Active Interventions. arXiv preprint, 2021.
- Interventions, Where and How? Experimental Design for Causal Models at Scale. Neural Information Processing Systems, 2022.
- Active Bayesian Causal Inference. Neural Information Processing Systems, 2022.
- Christopher M Bishop et al. Neural networks for pattern recognition. Oxford university press, 1995.
- Trajectory balance: Improved credit assignment in gflownets. arXiv preprint arXiv:2201.13259, 2022.
- Learning gflownets from partial episodes for improved convergence and stability. arXiv preprint arXiv:2209.12782, 2022.
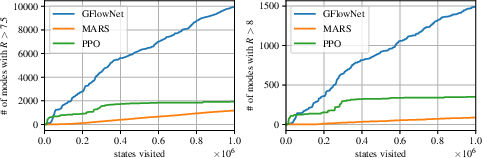
- {MARS}: Markov molecular sampling for multi-objective drug discovery. In International Conference on Learning Representations, 2021. URL https://openreview.net/forum?id=kHSu4ebxFXY.
- Biological sequence design with gflownets. In International Conference on Machine Learning, pages 9786–9801. PMLR, 2022a.
- Auto-encoding variational Bayes. International Conference on Learning Representations (ICLR), 2014.
- Stochastic backpropagation and approximate inference in deep generative models. International Conference on Machine Learning (ICML), 2014.
- Generative adversarial nets. In NIPS, 2014.
- A theory of continuous generative flow networks, 2023. URL https://arxiv.org/abs/2301.12594.
- Better training of gflownets with local credit and incomplete trajectories, 2023. URL https://arxiv.org/abs/2302.01687.
- Molecular mechanics-driven graph neural network with multiplex graph for molecular structures. arXiv preprint arXiv:2011.07457, 2020.
- The soluble epoxide hydrolase as a pharmaceutical target for hypertension. Journal of cardiovascular pharmacology, 50(3):225–237, 2007.
- Soluble epoxide hydrolase as a therapeutic target for cardiovascular diseases. Nature reviews Drug discovery, 8(10):794–805, 2009.
- Autodock vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. Journal of computational chemistry, 31(2):455–461, 2010.
- Zinc- a free database of commercially available compounds for virtual screening. Journal of chemical information and modeling, 45(1):177–182, 2005.
- Evaluating generalization in GFlownets for molecule design. In ICLR2022 Machine Learning for Drug Discovery, 2022. URL https://openreview.net/forum?id=JFSaHKNZ35b.
- Multi-objective gflownets. arXiv preprint arXiv:2210.12765, 2022b.
- Matthias Ehrgott. Multicriteria optimization, volume 491. Springer Science & Business Media, 2005.
- O’Neill UK government study. Antimicrobial resistance: Tackling a crisis for the health and wealth of nations, 2014.
- Bayesian structure learning with generative flow networks. In The 38th Conference on Uncertainty in Artificial Intelligence, 2022.
- Learning bayesian networks: The combination of knowledge and statistical data. Machine learning, 1995.
- Bayesian learning of causal structure and mechanisms with gflownets and variational bayes. arXiv preprint arXiv:2211.02763, 2022.
- Systematic discovery and perturbation of regulatory genes in human t cells reveals the architecture of immune networks. Nature Genetics, 54(8):1133–1144, 2022.
- Learning linear cyclic causal models with latent variables. The Journal of Machine Learning Research, 13(1):3387–3439, 2012.
- Amortized inference for causal structure learning. arXiv preprint arXiv:2205.12934, 2022b.
- Nodags-flow: Nonlinear cyclic causal structure learning. arXiv preprint arXiv:2301.01849, 2023.
- Bayesian Model Averaging. In Proceedings of the AAAI Workshop on Integrating Multiple Learned Models, 1996.
- Bayesian model averaging: a tutorial (with comments by M. Clyde, David Draper and EI George, and a rejoinder by the authors. Statistical science, 1999.
- Conditional neural processes. In International Conference on Machine Learning, pages 1704–1713. PMLR, 2018.
- Bayesian structure learning using dynamic programming and MCMC. Uncertainty in Artificial Intelligence, 2007.
- Sequential bayesian experimental design with variable cost structure. Advances in Neural Information Processing Systems, 33:4127–4137, 2020.
- Gflowout: Dropout with generative flow networks. arXiv preprint arXiv:2210.12928, 2022.
Paper Prompts
Sign up for free to create and run prompts on this paper using GPT-5.
Top Community Prompts
Collections
Sign up for free to add this paper to one or more collections.