- The paper introduces a triadic taxonomy categorizing ML applications based on the availability of governing equations.
- It details methodologies like supervised, reinforcement, and generative learning to tackle complex, multiscale scientific challenges.
- The work emphasizes developing interpretable, hybrid ML models to enable robust, generalizable, and causally-informed scientific insights.
Machine Learning as a Driver for Scientific Discovery: A Complexity-Based Taxonomy
Introduction
The reviewed paper presents a systematic framework for how ML methodologies can enable scientific discovery across disciplines—in particular, physics and the life sciences—by mapping ML approaches onto the degree of a priori knowledge available about the underlying system. The authors introduce a triadic taxonomy: problems with complete knowledge of governing equations, problems with partial knowledge, and problems with no tractable governing equations. This structure provides the scaffolding for understanding both successes and limitations of current ML-driven scientific advances, and for identifying open research frontiers.
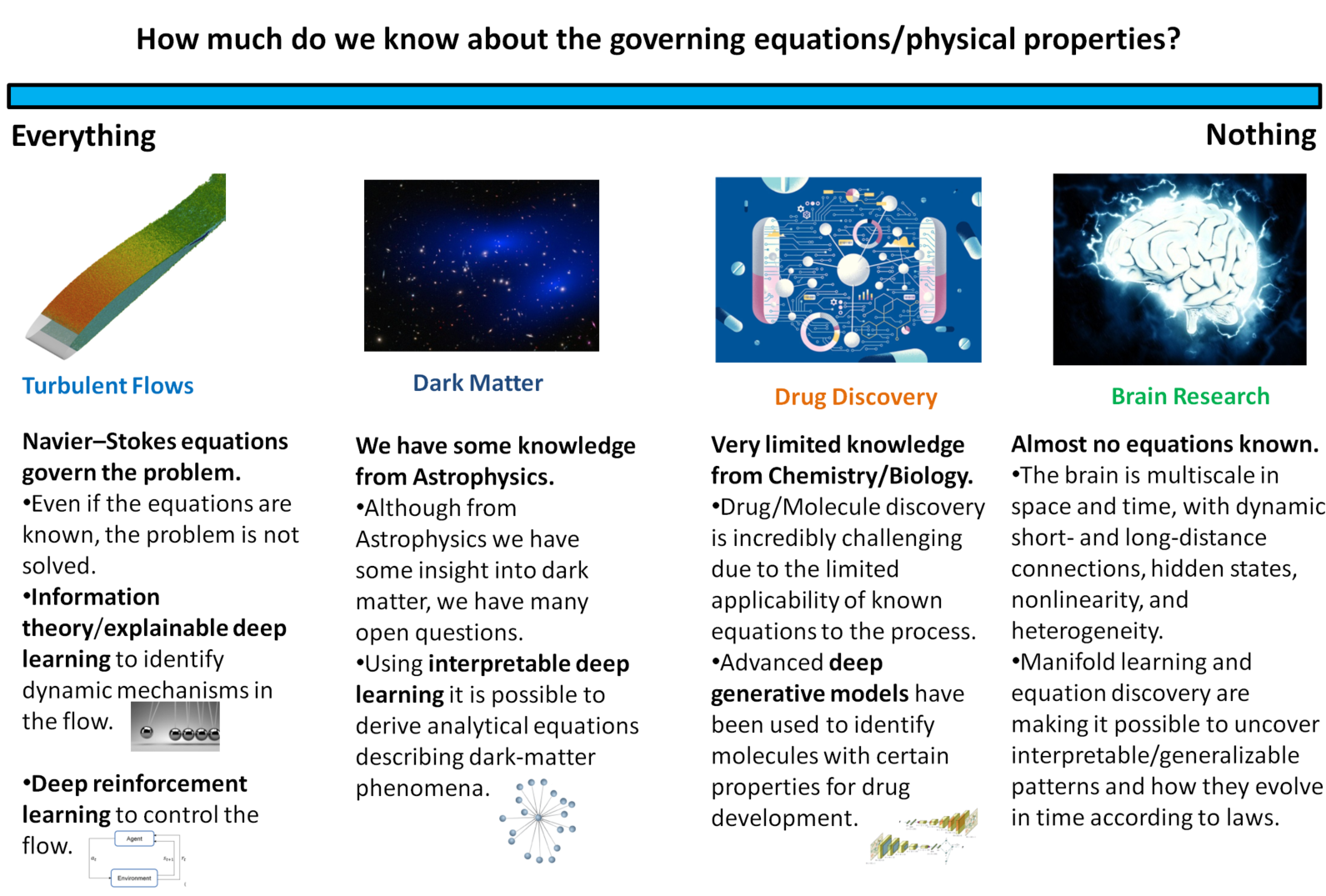

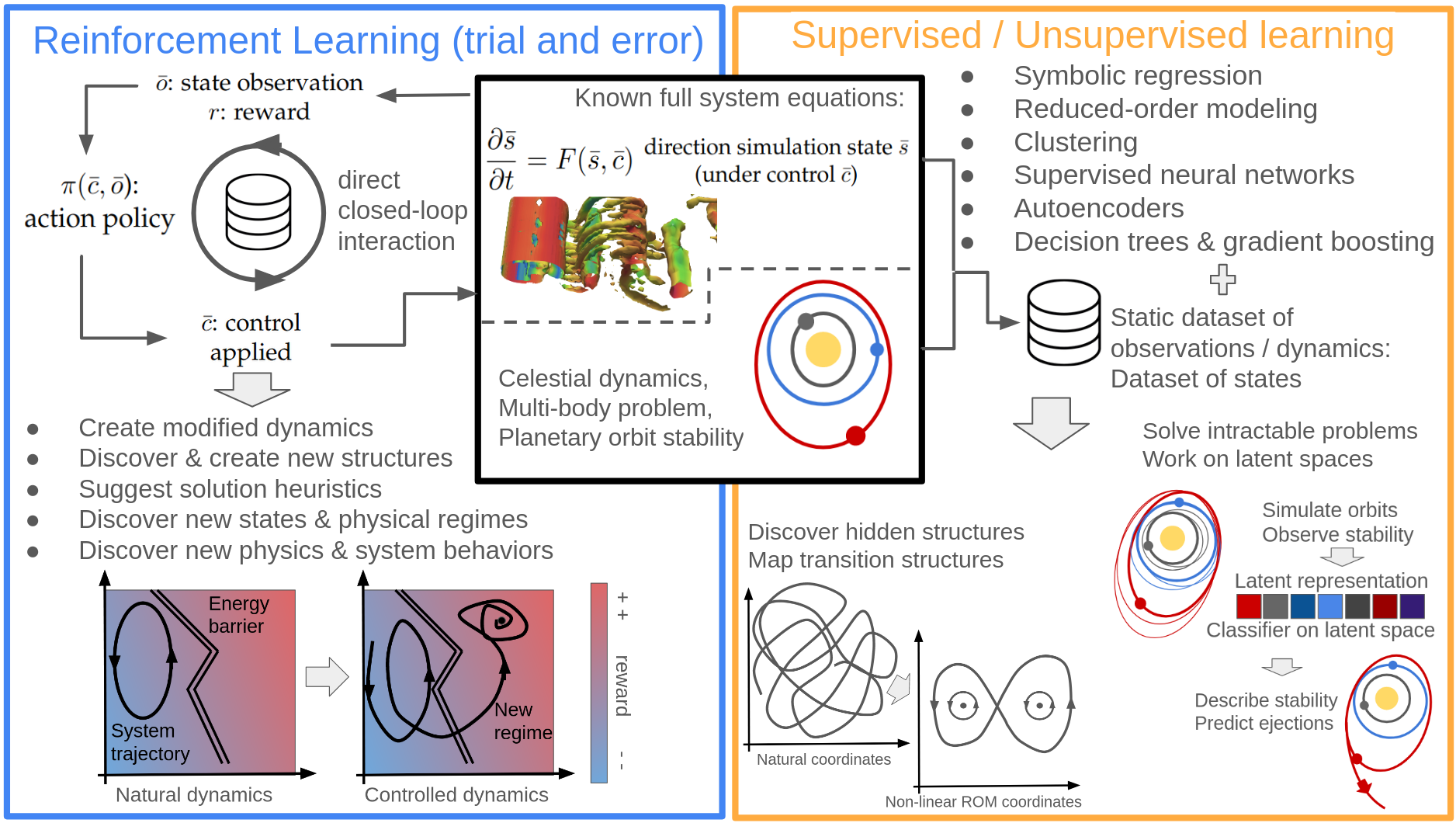
Figure 1: Schematic representation of the various applications of ML for scientific discovery, depending on the amount of knowledge available in each category.
ML When Governing Equations Are Known
In scientific domains such as turbulence, climate modeling, quantum physics, and certain astrophysical contexts, the governing equations are known in principle but direct simulation or analytic understanding remains non-trivial due to system complexity, multiscale interactions, or computational intractability. The review highlights several key ML strategies:
Most notably, the review points out that ML-driven approaches have begun to surpass classical methods not only in predictive accuracy but in their ability to generalize outside the classical training regime (e.g., SPOCK generalizes stability criteria in multi-planet systems beyond training distributions), with ML-derived surrogate models and fast simulators propelling discovery in domains ranging from particle physics to paleoclimate modeling.
ML with Partial Knowledge of Governing Equations
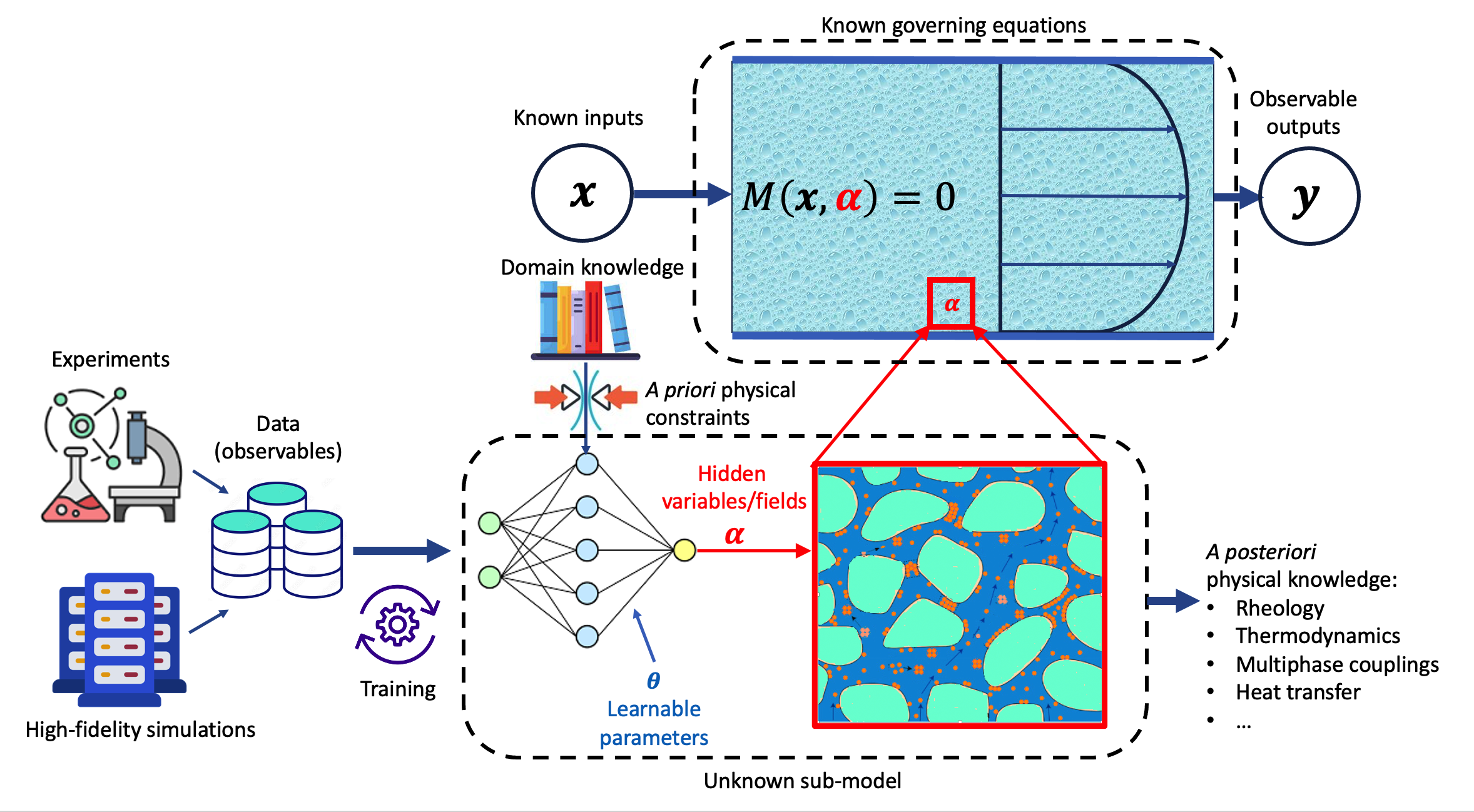
In intermediate regimes where only partial physical knowledge is available—such as in biochemistry, complex rheological flows, and multiscale or stochastic systems—the paper emphasizes hybrid modeling techniques:
The paper underscores a subtle, but important, epistemological point: These methods not only reproduce known phenomena but, in several cases, enable extrapolation—such as the transferability of protein backbones or discovery of functional motifs beyond training scenarios.
ML for Systems with No Known Governing Equations
In domains like neuroscience, certain areas of systems biology, or social science, where no closed-form governing equations exist, ML's role pivots to unsupervised discovery, latent structure modeling, and causal inference:
- Empirical Models and Representation Learning: Data-driven models (e.g., RNNs, NODEs) can be trained to emulate input-output relationships, recover latent dynamical manifolds, or predict experimental perturbations. Here, feature disentanglement and the use of interventional data are critical for extracting testable, mechanistic hypotheses.
(Figure 4)
Figure 4: Schematic representation of a model where the observed behavior depends on an unknown dynamic or causal structure; representation-learning methods can be employed to distil out underlying explanations.
- Causal Discovery and Invariant Structural Modeling: Emerging work on SINDy, CITRIS, and related methods show how sparse or piecewise ODE models can be inferred from data, sometimes uncovering interpretable mechanisms not accessible to classical statistical approaches.
- Scaling Challenges and Hybrid Approaches: The curse of dimensionality, incomplete measurements, and confounding variables present formidable challenges. The review highlights that scalable, active, closed-loop experimentation (often enabled by ML-driven imputation or dimensionality reduction) is necessary to make tractable progress in domains with high noise and partial observability.
- Cross-Pollination with ML Methodology: Diffusion models, flow-based generative models, and advances in self-supervised learning from ML are influencing scientific modeling both as tools for structure discovery and as objects of theoretical analysis, stimulating new directions in both fields.
Theoretical and Practical Implications
The paper makes several important claims:
- Interpretability as a Critical Bottleneck: Black-box approaches, despite predictive performance, remain epistemologically unsatisfying for scientific discovery unless interpretable surrogates, explainable AI, or symbolic regression are incorporated. Only this can ground claims of mechanism or causality.
- Generalization Beyond Training Distributions: The capacity of ML to identify transferable laws or motifs—across orders of magnitude in parameter regimes or molecular space—represents a distinct opportunity, but also introduces new risks regarding validity and extrapolation, demanding rigorous benchmarking and ablation within each scientific context.
- Validation Challenges: In the absence of ground-truth governing principles, validation of ML-derived discoveries must follow the classical scientific method, including iterative hypothesis testing, experiment design, and cross-domain replication.
- Data Scarcity and Simulation as a Solution: The model-driven generation of synthetic data, or the replacement of resource-intensive experiments with ML surrogates, can partially ameliorate data limitations, but introduces a dependency on simulation fidelity and training bias.
Outlook and Future Directions
Looking forward, the reviewed work posits several directions for AI-assisted scientific discovery:
- Development of Generalizable Hybrid Models: Integrating symbolic reasoning, physics priors, and data-driven learning is essential to move from mere prediction to explanation and hypothesis generation, particularly via architectures capable of embedding known constraints or symmetries.
- Scalable, Interpretable ML Pipelines for Discovery: The demand for explainable and interpretable methodologies will grow, particularly in high-impact domains (biomedicine, climate modeling), necessitating new methods for model introspection and uncertainty quantification.
- Causal and Mechanistic Inference Beyond Correlation: There is growing recognition of the necessity of formal causal inference frameworks—encompassing interventions, invariances, and representation learning—for progress in fields where data is observational or experimental manipulation is feasible.
- Cross-Disciplinary and Meta-Scientific Opportunities: ML serves as both a tool for accelerating discovery and as an object of scientific inquiry itself, e.g., in the design of new algorithms, combinatorial optimization, or meta-learning, thereby generating a virtuous cycle between scientific practice and ML research.
Conclusion
The paper provides a nuanced and careful synthesis of how ML is currently redefining the landscape of scientific discovery. The taxonomy rooted in the degree of prior knowledge enables targeted discussions of appropriate ML methods, their interpretability, and their validation challenges. While ML enables unprecedented progress in extracting insight from complexity—ranging from analytically intractable PDEs to neural circuits—the critical path forward remains tied to improving interpretability, ground-truth validation, and hybrid models that robustly encode domain knowledge. Future developments in AI for science will likely hinge on advances that unify explainability, out-of-distribution robustness, and scalable generalization.