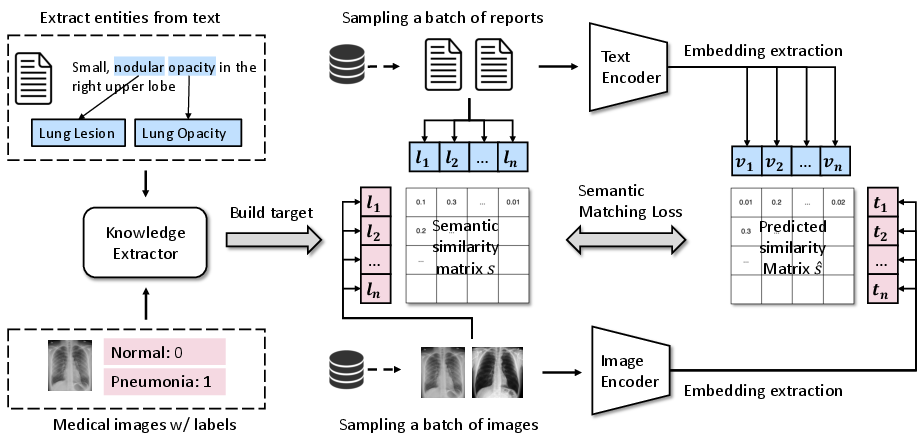
- The paper introduces a decoupled contrastive learning framework that replaces InfoNCE with a semantic matching loss to tackle unpaired medical datasets.
- It leverages state-of-the-art vision and text encoders, including Swin Transformer and BioClinicalBERT, for effective multimodal semantic alignment.
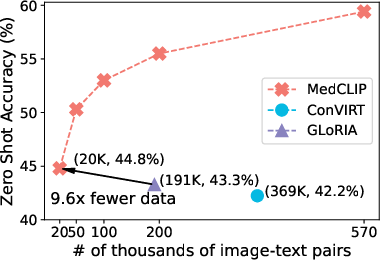
- Experiments on datasets like CheXpert and MIMIC-CXR show over a 10% accuracy improvement in zero-shot classification using only 10% of the data.
MedCLIP: Contrastive Learning from Unpaired Medical Images and Text
Introduction
The paper "MedCLIP: Contrastive Learning from Unpaired Medical Images and Text" addresses the challenges of using contrastive learning in the medical domain, where the availability of paired image-text data is significantly limited compared to general-domain datasets. Traditional methods like CLIP are limited by their dependence on large amounts of such paired data. In contrast, MedCLIP introduces a novel approach that decouples images and texts, leveraging combinatorial data expansion and medical semantics to improve data efficiency and model performance.
Challenges and Novel Approaches
Medical image-text datasets often suffer from insufficient data and the presence of false negatives. This paper identifies key challenges:
- Data Insufficiency: The scarcity of paired medical image-text data poses a significant hurdle. MedCLIP effectively addresses this by decoupling images and texts to utilize vast unpaired datasets, scaling the training data to a combinatorial magnitude.
- Semantic Mismatch and False Negatives: Traditional methods like InfoNCE introduce false negatives by misclassifying semantically related but unpaired data as negatives. MedCLIP overcomes this by replacing InfoNCE with a semantic matching loss, grounded in medical domain knowledge, to align image and text semantics accurately.
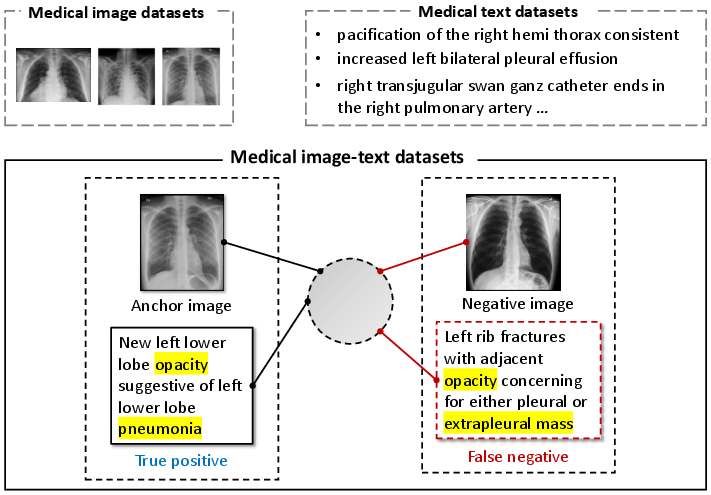
Figure 1: Illustration of challenges in medical image-text contrastive learning such as ignored datasets and false negatives.
Methodology
MedCLIP's architecture consists of a vision and text encoder system that utilizes domain-specific knowledge for semantic alignment between data sources.
Experiments and Results
Extensive experimentation across multiple datasets (CheXpert, MIMIC-CXR) demonstrates MedCLIP's superiority in several key metrics:
Implications and Future Directions
MedCLIP’s approach facilitates broader application of multimodal learning in the medical domain, showcasing notable improvements in data efficiency and semantic understanding. By effectively harnessing unpaired datasets and medical knowledge, MedCLIP sets a foundation for future innovations in medical AI, suggesting potential for further exploration in automated diagnosis and cross-domain knowledge applications.
Conclusion
MedCLIP successfully introduces a framework for decoupled contrastive learning, overcoming traditional limitations in medical image-text pre-training. Its innovative use of domain semantics and efficient data utilization positions MedCLIP as a strategic advancement in medical AI, facilitating more semantic-rich and resource-efficient model training. Future works could optimize semantic processing and further expand on this foundational work.