- The paper introduces a novel integration of genome-scale metabolic modeling and ecological network analysis to predict probiotic engraftment in gut microbiota.
- It employs pairwise flux balance analysis and dynamical models, including modified Lotka-Volterra, to capture microbial interactions with high predictive accuracy.
- The findings suggest that mechanistic modeling can outperform traditional machine learning, supporting personalized probiotic design in complex microbial communities.
Introduction
The paper "Metabolic Model-based Ecological Modeling for Probiotic Design" (2210.03198) addresses the challenge of predicting the engraftment potential of probiotics within the human gut microbiota. The study leverages genome-scale metabolic modeling (GSMM) and network analysis to anticipate which bacterial strains can successfully integrate into existing microbial communities. The approach is based on the hypothesis that biotic interactions, mediated by metabolic exchanges, significantly influence the success of these microbial introductions.
Methodology
The study utilizes genome-scale metabolic models (GEMs) available in the AGORA database, which includes 818 bacterial species. Pairwise metabolic modeling is conducted to generate interaction networks, scored by six dynamical systems models, including the generalized Lotka-Volterra (LV) model and its variants. The LV model is known for its ability to capture population dynamics, despite its dependency on extensive time-longitudinal data for parameterization—a limitation addressed by integrating GSMM-derived constraints.
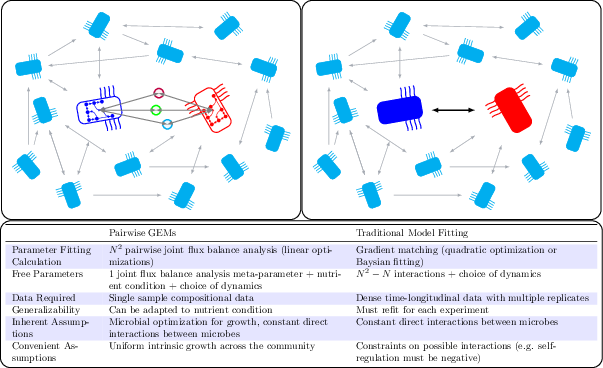
Figure 1: Population models such as Lotka-Volterra generally require dense time-longitudinal data to accurately parameterize. In this work, we leverage genome-scale metabolic modeling to parameterize population models with only genomic data from a single time-point. We are unable to parameterize these models using standard techniques due to the sparsity of the data.
The core methodology involves constructing interaction networks using joint flux balance analysis on pairs of species, transforming raw metabolic data into interaction coefficients that inform dynamic models predicting engraftment likelihood in different microbial communities. This is validated across three different empirical datasets involving both probiotic bacteria and pathogenic species.
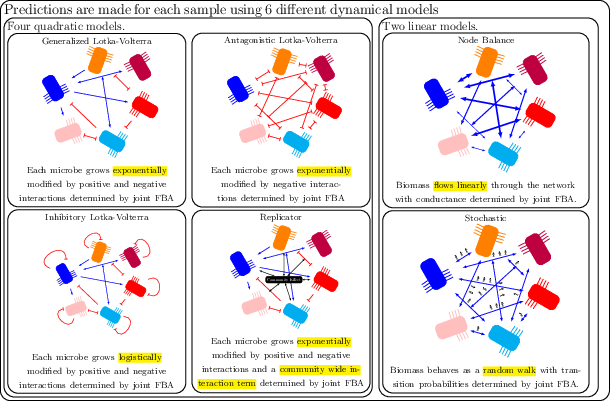
Figure 2: We used six dynamical systems to assess the engraftment potential of an invading taxa. All six are parameterized based on a set of network connections wij derived from metabolic modeling.
Results
The results demonstrate the model's efficacy in predicting microbial engraftment, surpassing traditional machine learning techniques like support vector machines and random forests in scenarios with limited data or high community complexity. Notably, the modified antagonistic and inhibitory versions of the Lotka-Volterra model prevented numerical instabilities while delivering robust predictions.
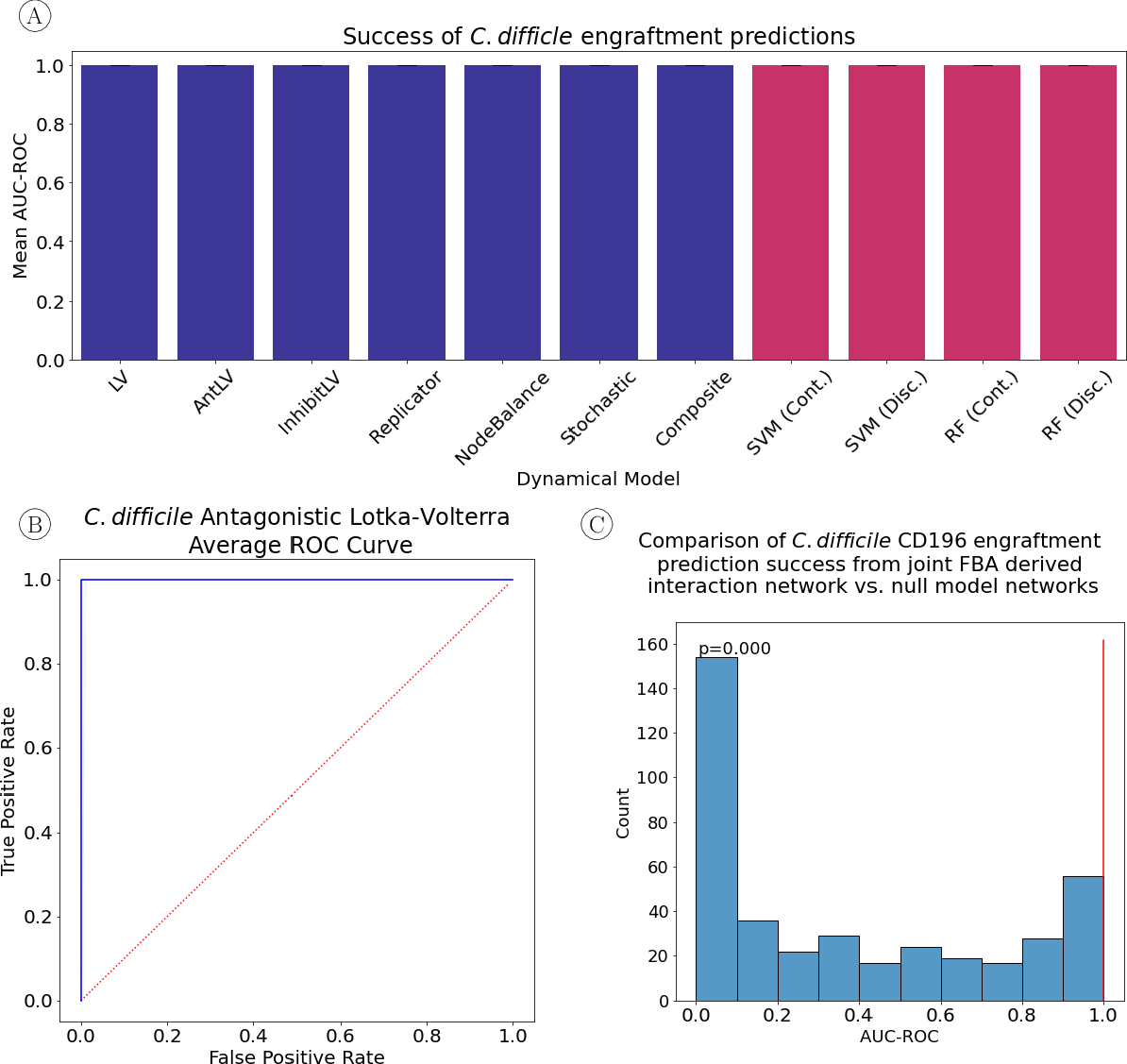
Figure 3: {\bf (A)} All network dynamics tested accurately predicted the engraftment versus non-engraftment as determined by the mean area under the receiver operator characteristic curve (AUC-ROC).
In the case of Clostridioides difficile, the model achieved perfect classification, indicating its potential utility in distinguishing between susceptible and resistant microbial profiles. With probiotic candidates Bifidobacterium longum and Lactiplantibacillus plantarum, the system's performance varied, yet consistently outperformed non-mechanistic approaches.
Discussion
The paper posits that predicting microbial engraftment using metabolic interaction networks offers a pathway toward more personalized probiotic therapies. This approach enhances understanding of microbial community dynamics, central to therapeutic strategies aiming to modulate gut health. The mechanistic nature of the study reveals not just interaction patterns but also insights into microbial competition and cooperation mechanisms, including the potential for synthetic ecological designs to optimize probiotic efficacy.
Limitations regarding the fidelity of network models in entirely new environments and the potential discrepancies in metabolic network reconstructions indicate areas for further research. Enhancing environmental adaptability and expanding databases to include more comprehensive GEMs could address these constraints.
Conclusion
The integration of genome-scale metabolic modeling and ecological network analysis presents a promising method for predicting and enhancing probiotic engraftment in gut ecosystems. The results demonstrate that such an approach holds promise for designing effective microbiome-based interventions, laying a foundation for future work aimed at refining dynamical models to enhance their precision and applicability in diverse microbial contexts. Further development will focus on augmenting model adaptability and robustness, potentially incorporating real-time environmental feedback to predict and alter microbial compositions dynamically.




